Accelerate Correlation with GPUs
This example shows how to use a GPU to accelerate cross-correlation. Many correlation problems involve large data sets and can be solved much faster using a GPU. This example requires a Parallel Computing Toolbox™ user license. Refer to GPU Computing Requirements (Parallel Computing Toolbox) to see what GPUs are supported.
Introduction
Start by learning some basic information about the GPU in your machine. To access the GPU, use the Parallel Computing Toolbox.
if ~(parallel.gpu.GPUDevice.isAvailable) fprintf("\n\t**GPU not available. Stopping.**\n"); return; else dev = gpuDevice; fprintf(... "GPU detected (%s, %d multiprocessors, Compute Capability %s)", ... dev.Name, dev.MultiprocessorCount, dev.ComputeCapability); end
GPU detected (NVIDIA RTX A5000, 64 multiprocessors, Compute Capability 8.6)
Benchmarking Functions
Because code written for the CPU can be ported to run on the GPU, a single function can be used to benchmark both the CPU and GPU. However, because code on the GPU executes asynchronously from the CPU, special precaution should be taken when measuring performance. Before measuring the time taken to execute a function, ensure that all GPU processing has finished by executing the 'wait' method on the device. This extra call will have no effect on the CPU performance.
This example benchmarks three different types of cross-correlation.
Benchmark Simple Cross-Correlation
For the first case, two vectors of equal size are cross-correlated using the syntax xcorr(u,v). The ratio of CPU execution time to GPU execution time is plotted against the size of the vectors.
sizes = floor(logspace(5,7,10))'; tc = zeros(numel(sizes),1); tg = zeros(numel(sizes),1); for s = 1:numel(sizes) if s == 1 fprintf("** Benchmarking vector-vector cross-correlation. **") fprintf("Running xcorr of N elements") end fprintf("."); a = single(rand(sizes(s),1)); b = single(rand(sizes(s),1)); tc(s) = timeit(@() xcorr(a,b),1); tg(s) = gputimeit(@() xcorr(gpuArray(a),gpuArray(b)),1); if s == numel(sizes) disp(table(sizes,1e3*tc,1e3*tg,VariableNames= ... ["Length (N)" "CPU time (ms)" "GPU time (ms)"])) end end
** Benchmarking vector-vector cross-correlation. **
Running xcorr of N elements
..........
Length (N) CPU time (ms) GPU time (ms)
__________ _____________ _____________
1e+05 3.2248 1.1345
1.6681e+05 6.0368 1.2935
2.7826e+05 9.0778 1.5205
4.6416e+05 16.32 1.9185
7.7426e+05 25.602 2.5235
1.2915e+06 56.349 3.7115
2.1544e+06 85.022 7.5365
3.5938e+06 143.4 13.051
5.9948e+06 285.56 20.219
1e+07 365.81 33.806
% Plot the results fig = figure; ax = axes(Parent=fig); semilogx(ax,sizes,tc./tg,"r*-"); ylabel(ax,"Speedup"); xlabel(ax,"Vector size"); title(ax,"GPU Acceleration of XCORR");

Benchmarking Matrix Column Cross-Correlation
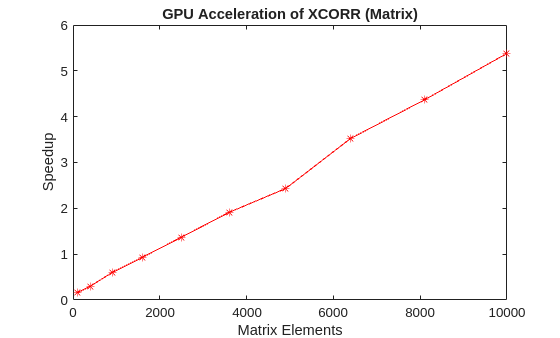
For the second case, the columns of a matrix A are pairwise cross-correlated to produce a large matrix output of all correlations using the syntax xcorr(A). The ratio of CPU execution time to GPU execution time is plotted against the size of the matrix A.
sizes = floor(logspace(2,2.5,10))'; tc = zeros(numel(sizes),1); tg = zeros(numel(sizes),1); for s = 1:numel(sizes) if s == 1 fprintf("** Benchmarking matrix column cross-correlation. **") fprintf("Running xcorr (matrix) of an N-by-N matrix") end fprintf("."); a = single(rand(sizes(s))); tc(s) = timeit(@() xcorr(a),1); tg(s) = gputimeit(@() xcorr(gpuArray(a)),1); if s == numel(sizes) disp(table(sizes,1e3*tc,1e3*tg,VariableNames= ... ["Size (N)" "CPU time (ms)" "GPU time (ms)"])) end end
** Benchmarking matrix column cross-correlation. **
Running xcorr (matrix) of an N-by-N matrix
..........
Size (N) CPU time (ms) GPU time (ms)
________ _____________ _____________
100 5.3818 0.80717
113 8.0898 0.88967
129 13.426 1.6885
146 18.493 2.0345
166 25.6 2.4435
189 36.405 3.1765
215 57.449 3.8975
244 81.755 4.7185
278 131.18 11.829
316 178.32 15.023
% Plot the results fig = figure; ax = axes(Parent=fig); semilogx(ax,sizes.^2,tc./tg,"r*-") ylabel(ax,"Speedup") xlabel(ax,"Matrix Elements") title(ax,"GPU Acceleration of XCORR (Matrix)")
Benchmarking Two-Dimensional Cross-Correlation
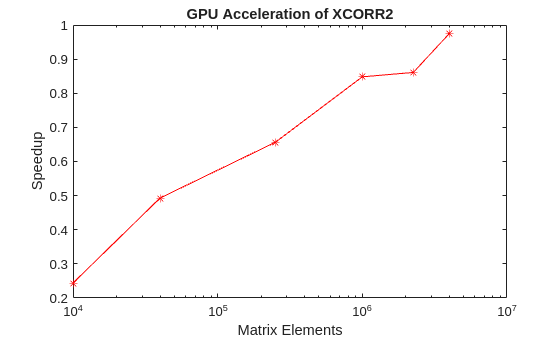
For the final case, two matrices, X and Y, are cross correlated using xcorr2(X,Y). X is fixed in size while Y is allowed to vary. The speedup is plotted against the size of the second matrix.
sizes = floor(logspace(2.5,3.5,10))'; tc = zeros(numel(sizes),1); tg = zeros(numel(sizes),1); a = single(rand(100)); for s = 1:numel(sizes) if s == 1 fprintf("** Benchmarking 2-D cross-correlation**") fprintf("Running xcorr2 of 100-by-100 and N-by-N matrices.") end fprintf("."); b = single(rand(sizes(s))); tc(s) = timeit(@() xcorr2(a,b),1); tg(s) = gputimeit(@() xcorr2(gpuArray(a),gpuArray(b)),1); if s == numel(sizes) disp(table(sizes,1e3*tc,1e3*tg,VariableNames= ... ["Size (N)" "CPU time (ms)" "GPU time (ms)"])) end end
** Benchmarking 2-D cross-correlation**
Running xcorr2 of 100-by-100 and N-by-N matrices.
..........
Size (N) CPU time (ms) GPU time (ms)
________ _____________ _____________
316 3.9288 2.7275
408 6.0308 2.9065
527 9.4288 3.4885
681 15.166 5.8805
879 24.238 7.5805
1136 37.586 12.254
1467 65.472 18.775
1895 109.37 28.703
2448 171.28 42.79
3162 299.2 70.185
% Plot the results fig = figure; ax =axes(Parent=fig); semilogx(ax,sizes.^2,tc./tg,"r*-") ylabel(ax,"Speedup") xlabel(ax,"Matrix Elements") title(ax,"GPU Acceleration of XCORR2")
Other GPU Accelerated Signal Processing Functions
There are several other signal processing functions that can be run on the GPU. These functions include fft, ifft, conv, filter, fftfilt, and more. In some cases, you can achieve large acceleration relative to the CPU. For a full list of GPU accelerated signal processing functions, see the GPU Algorithm Acceleration section in the Signal Processing Toolbox™ documentation.
See Also
gather (Parallel Computing Toolbox) | gpuArray (Parallel Computing Toolbox) | xcorr