Plot the prey versus predator data from the stochastically simulated lotka model in separate subplots by using a custom function (plotXY).
Load the model. Set the solver type to SSA to perform stochastic simulations, and set the stop time to 3.
Set the number of runs and use sbioensemblerun for simulation.
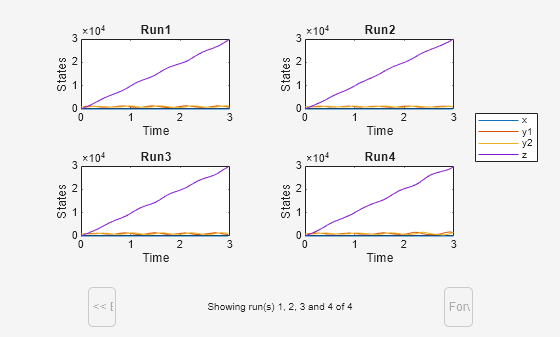
Plot each simulation run in a separate subplot. By default, sbiosubplot shows the time plot of each species for each run per subplot.
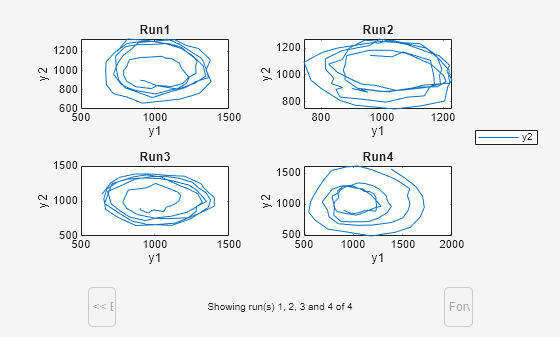
Plot selected states against each other; in this case, plot the prey population versus the predator population in separate subplots for each run. Use the function plotXY (shown at the end of this example) to plot the simulated y1 (prey) data versus the y2 (predator). Specify the function as a function handle in the sbiosubplot call to plot each run in its own subplot. In this case, the fifth input argument (showLegend) is set to true, which means the fourth input argument (yArgs) is shown as the legend.
If you use the live script file for this example, the plotXY function is already included at the end of the file. Otherwise, you must define the plotXY function at the end of your .m or .mlx file or add it as a file on the MATLAB path.
Define plotXY Function
sbiosubplot accepts a function handle of a function with the signature:
function functionName(sd,xArgs,yArgs).
The plotXY function plots two selected states against each other. The first input sd is the simulation data (SimBiology SimData object or vector of objects). In this example, xArgs is a cell array containing the name of the species to be plotted on the x-axis, and yArgs is a cell array containing the name of the species to be plotted on the y-axis. However, you can use the inputs xArgs and yArgs in any way in your custom plotting function. No output from the function is necessary.