fitcknn
Fit k-nearest neighbor classifier
Syntax
Description
Mdl = fitcknn(Tbl,ResponseVarName)Tbl and output (response)
Tbl.ResponseVarName.
Mdl = fitcknn(___,Name=Value)
[
also returns Mdl,AggregateOptimizationResults] = fitcknn(___)AggregateOptimizationResults, which contains
hyperparameter optimization results when you specify the
OptimizeHyperparameters and
HyperparameterOptimizationOptions name-value arguments.
You must also specify the ConstraintType and
ConstraintBounds options of
HyperparameterOptimizationOptions. You can use this
syntax to optimize on compact model size instead of cross-validation loss, and
to perform a set of multiple optimization problems that have the same options
but different constraint bounds.
Examples
Train a k-nearest neighbor classifier using Fisher's iris data, where k, the number of nearest neighbors in the predictors, is 5.
Load Fisher's iris data.
load fisheriris
X = meas;
Y = species;X is a numeric matrix that contains four measurements for 150 irises. Y is a cell array of character vectors that contains the corresponding iris species.
Train a 5-nearest neighbor classifier. Standardize the noncategorical predictor data.
Mdl = fitcknn(X,Y,NumNeighbors=5,Standardize=true)
Mdl =
ClassificationKNN
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'setosa' 'versicolor' 'virginica'}
ScoreTransform: 'none'
NumObservations: 150
Distance: 'euclidean'
NumNeighbors: 5
Properties, Methods
Mdl is a trained ClassificationKNN classifier.
To access the properties of Mdl, use dot notation.
Mdl.ClassNames
ans = 3×1 cell
{'setosa' }
{'versicolor'}
{'virginica' }
Mdl.Prior
ans = 1×3
0.3333 0.3333 0.3333
Mdl.Prior contains the class prior probabilities, which you can specify using the Prior name-value argument in fitcknn. The order of the class prior probabilities corresponds to the order of the classes in Mdl.ClassNames. By default, the prior probabilities are the respective relative frequencies of the classes in the data.
You can also reset the prior probabilities after training. For example, set the prior probabilities to 0.5, 0.2, and 0.3, respectively.
Mdl.Prior = [0.5 0.2 0.3];
You can pass Mdl to predict to label new measurements or crossval to cross-validate the classifier.
Train a k-nearest neighbor classifier using the Minkowski distance metric.
Load Fisher's iris data set.
load fisheriris
X = meas;
Y = species;X is a numeric matrix that contains four measurements for 150 irises. Y is a cell array of character vectors that contains the corresponding iris species.
Train a 3-nearest neighbors classifier using the Minkowski metric. To use the Minkowski metric, you must use an exhaustive searcher. Standardize the noncategorical predictor data.
Mdl = fitcknn(X,Y,NumNeighbors=3, ... NSMethod="exhaustive",Distance="minkowski", ... Standardize=true);
Mdl is a ClassificationKNN classifier.
Find the Minkowski distance exponent used to train Mdl.
Mdl.DistParameter
ans = 2
You can modify a distance parameter after creating a ClassificationKNN object. For example, set the Minkowski distance exponent to 4.
Mdl.DistParameter = 4; Mdl.DistParameter
ans = 4
Train a k-nearest neighbor classifier using the chi-square distance.
Load Fisher's iris data set.
load fisheriris
X = meas;
Y = species;The chi-square distance between j-dimensional points x and z is
where is a weight associated with dimension j.
Specify the chi-square distance function. The distance function must:
Take one row of
X, for example,x, and the matrixZ.Compare
xto each row ofZ.Return a vector
Dof length , where is the number of rows ofZ. Each element ofDis the distance between the observation corresponding toxand the observations corresponding to each row ofZ.
chiSqrDist = @(x,Z,wt)sqrt(((x-Z).^2)*wt);
This example uses arbitrary weights for illustration.
Train a 3-nearest neighbor classifier. It is good practice to standardize noncategorical predictor data.
k = 3;
w = [0.3; 0.3; 0.2; 0.2];
KNNMdl = fitcknn(X,Y,Distance=@(x,Z)chiSqrDist(x,Z,w), ...
NumNeighbors=k,Standardize=true);KNNMdl is a ClassificationKNN classifier.
Cross validate the KNN classifier using the default 10-fold cross validation. Examine the classification error.
rng(1); % For reproducibility
CVKNNMdl = crossval(KNNMdl);
classError = kfoldLoss(CVKNNMdl)classError = 0.0600
CVKNNMdl is a ClassificationPartitionedModel classifier.
Compare the classifier with one that uses a different weighting scheme.
w2 = [0.2; 0.2; 0.3; 0.3];
CVKNNMdl2 = fitcknn(X,Y,Distance=@(x,Z)chiSqrDist(x,Z,w2), ...
NumNeighbors=k,KFold=10,Standardize=true);
classError2 = kfoldLoss(CVKNNMdl2)classError2 = 0.0400
The second weighting scheme yields a classifier that has better out-of-sample performance.
Automatically optimize hyperparameters of a k-nearest neighbor classifier by using fitcknn.
Load the fisheriris data set.
load fisheriris
X = meas;
Y = species;Find hyperparameters that minimize the 5-fold cross-validation loss by using automatic hyperparameter optimization. For reproducibility, set the random seed and use the "expected-improvement-plus" acquisition function.
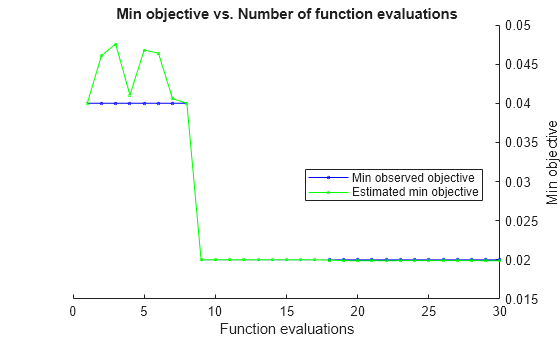
rng(1) hpoOptions = hyperparameterOptimizationOptions(AcquisitionFunctionName="expected-improvement-plus"); Mdl = fitcknn(X,Y,OptimizeHyperparameters="auto", ... HyperparameterOptimizationOptions=hpoOptions)
|====================================================================================================================|
| Iter | Eval | Objective | Objective | BestSoFar | BestSoFar | NumNeighbors | Distance | Standardize |
| | result | | runtime | (observed) | (estim.) | | | |
|====================================================================================================================|
| 1 | Best | 0.04 | 0.41125 | 0.04 | 0.04 | 13 | minkowski | true |
| 2 | Accept | 0.19333 | 0.19876 | 0.04 | 0.046097 | 1 | correlation | true |
| 3 | Accept | 0.053333 | 0.053887 | 0.04 | 0.047573 | 14 | chebychev | true |
| 4 | Accept | 0.046667 | 0.048087 | 0.04 | 0.041053 | 2 | minkowski | false |
| 5 | Accept | 0.053333 | 0.057237 | 0.04 | 0.046782 | 7 | minkowski | true |
| 6 | Accept | 0.10667 | 0.095523 | 0.04 | 0.046422 | 2 | mahalanobis | false |
| 7 | Accept | 0.093333 | 0.036604 | 0.04 | 0.040581 | 75 | minkowski | false |
| 8 | Accept | 0.15333 | 0.026665 | 0.04 | 0.040008 | 75 | minkowski | true |
| 9 | Best | 0.02 | 0.025095 | 0.02 | 0.02001 | 4 | minkowski | false |
| 10 | Accept | 0.026667 | 0.028616 | 0.02 | 0.020012 | 8 | minkowski | false |
| 11 | Accept | 0.21333 | 0.045495 | 0.02 | 0.020008 | 69 | chebychev | true |
| 12 | Accept | 0.053333 | 0.041943 | 0.02 | 0.020009 | 5 | chebychev | true |
| 13 | Accept | 0.053333 | 0.041198 | 0.02 | 0.020009 | 1 | chebychev | true |
| 14 | Accept | 0.053333 | 0.063575 | 0.02 | 0.020008 | 5 | seuclidean | false |
| 15 | Accept | 0.053333 | 0.03088 | 0.02 | 0.020008 | 21 | seuclidean | false |
| 16 | Accept | 0.053333 | 0.035074 | 0.02 | 0.020009 | 1 | seuclidean | false |
| 17 | Accept | 0.15333 | 0.031856 | 0.02 | 0.020007 | 75 | seuclidean | false |
| 18 | Accept | 0.02 | 0.025901 | 0.02 | 0.019969 | 5 | minkowski | false |
| 19 | Accept | 0.33333 | 0.08659 | 0.02 | 0.019898 | 2 | spearman | false |
| 20 | Accept | 0.23333 | 0.03016 | 0.02 | 0.019888 | 71 | mahalanobis | false |
|====================================================================================================================|
| Iter | Eval | Objective | Objective | BestSoFar | BestSoFar | NumNeighbors | Distance | Standardize |
| | result | | runtime | (observed) | (estim.) | | | |
|====================================================================================================================|
| 21 | Accept | 0.046667 | 0.035697 | 0.02 | 0.019895 | 1 | cityblock | true |
| 22 | Accept | 0.053333 | 0.032859 | 0.02 | 0.019892 | 6 | cityblock | true |
| 23 | Accept | 0.12 | 0.036361 | 0.02 | 0.019895 | 75 | cityblock | true |
| 24 | Accept | 0.06 | 0.027765 | 0.02 | 0.019903 | 2 | cityblock | false |
| 25 | Accept | 0.033333 | 0.02355 | 0.02 | 0.019899 | 17 | cityblock | false |
| 26 | Accept | 0.12 | 0.024571 | 0.02 | 0.019907 | 74 | cityblock | false |
| 27 | Accept | 0.033333 | 0.024424 | 0.02 | 0.019894 | 7 | cityblock | false |
| 28 | Accept | 0.02 | 0.024401 | 0.02 | 0.019897 | 1 | chebychev | false |
| 29 | Accept | 0.02 | 0.02943 | 0.02 | 0.019891 | 4 | chebychev | false |
| 30 | Accept | 0.08 | 0.070035 | 0.02 | 0.019891 | 28 | chebychev | false |
__________________________________________________________
Optimization completed.
MaxObjectiveEvaluations of 30 reached.
Total function evaluations: 30
Total elapsed time: 10.5839 seconds
Total objective function evaluation time: 1.7435
Best observed feasible point:
NumNeighbors Distance Standardize
____________ _________ ___________
4 minkowski false
Observed objective function value = 0.02
Estimated objective function value = 0.020124
Function evaluation time = 0.025095
Best estimated feasible point (according to models):
NumNeighbors Distance Standardize
____________ _________ ___________
5 minkowski false
Estimated objective function value = 0.019891
Estimated function evaluation time = 0.0259
Mdl =
ClassificationKNN
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'setosa' 'versicolor' 'virginica'}
ScoreTransform: 'none'
NumObservations: 150
HyperparameterOptimizationResults: [1×1 classreg.learning.paramoptim.SupervisedLearningBayesianOptimization]
Distance: 'minkowski'
NumNeighbors: 5
Properties, Methods
The trained classifier Mdl corresponds to the best estimated feasible point and uses the same hyperparameter values for NumNeighbors, Distance, and Standardize.
Find the hyperparameter values used to train Mdl by using the bestPoint function. By default, bestPoint uses the same best point criterion used by fitcknn during the hyperparameter optimization ("min-visited-upper-confidence-interval"). In general, fit functions determine the best hyperparameter values based on the "min-visited-upper-confidence-interval" criterion (instead of the "min-observed" criterion) to avoid overfitting to noise in the data set.
bestEstimatedPoint = bestPoint(Mdl.HyperparameterOptimizationResults)
bestEstimatedPoint=1×3 table
NumNeighbors Distance Standardize
____________ _________ ___________
5 minkowski false
Verify that the results match the properties of Mdl. Note that the Mu and Sigma properties of a ClassificationKNN object are empty when the k-nearest neighbor classifier does not use standardization.
classifierProperties = table(Mdl.NumNeighbors,string(Mdl.Distance), ... struct(Means=Mdl.Mu,StandardDeviations=Mdl.Sigma), ... VariableNames=["NumNeighbors","Distance","Standardize"])
classifierProperties=1×3 table
NumNeighbors Distance Standardize
____________ ___________ ___________
5 "minkowski" 1×1 struct
classifierProperties.Standardize
ans = struct with fields:
Means: []
StandardDeviations: []
Input Arguments
Sample data used to train the model, specified as a table. Each row of Tbl
corresponds to one observation, and each column corresponds to one predictor variable.
Optionally, Tbl can contain one additional column for the response
variable. Multicolumn variables and cell arrays other than cell arrays of character
vectors are not allowed.
If
Tblcontains the response variable, and you want to use all remaining variables inTblas predictors, then specify the response variable by usingResponseVarName.If
Tblcontains the response variable, and you want to use only a subset of the remaining variables inTblas predictors, then specify a formula by usingformula.If
Tbldoes not contain the response variable, then specify a response variable by usingY. The length of the response variable and the number of rows inTblmust be equal.
Response variable name, specified as the name of a variable in
Tbl.
You must specify ResponseVarName as a character vector or string scalar.
For example, if the response variable Y is
stored as Tbl.Y, then specify it as
"Y". Otherwise, the software
treats all columns of Tbl, including
Y, as predictors when training
the model.
The response variable must be a categorical, character, or string array; a logical or numeric
vector; or a cell array of character vectors. If
Y is a character array, then each
element of the response variable must correspond to one row of
the array.
A good practice is to specify the order of the classes by using the
ClassNames name-value
argument.
Data Types: char | string
Explanatory model of the response variable and a subset of the predictor variables,
specified as a character vector or string scalar in the form
"Y~x1+x2+x3". In this form, Y represents the
response variable, and x1, x2, and
x3 represent the predictor variables.
To specify a subset of variables in Tbl as predictors for
training the model, use a formula. If you specify a formula, then the software does not
use any variables in Tbl that do not appear in
formula.
The variable names in the formula must be both variable names in Tbl
(Tbl.Properties.VariableNames) and valid MATLAB® identifiers. You can verify the variable names in Tbl by
using the isvarname function. If the variable names
are not valid, then you can convert them by using the matlab.lang.makeValidName function.
Data Types: char | string
Class labels, specified as a categorical, character, or string array, a logical or numeric
vector, or a cell array of character vectors. Each row of Y
represents the classification of the corresponding row of X.
The software considers NaN, '' (empty character vector),
"" (empty string), <missing>, and
<undefined> values in Y to be missing
values. Consequently, the software does not train using observations with a missing
response.
Data Types: categorical | char | string | logical | single | double | cell
Predictor data, specified as numeric matrix.
Each row corresponds to one observation (also known as an instance or example), and each column corresponds to one predictor variable (also known as a feature).
The length of Y and the number of rows of
X must be equal.
To specify the names of the predictors in the order of their appearance in
X, use the PredictorNames
name-value pair argument.
Data Types: double | single
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: fitcknn(Tbl,Y,NumNeighbors=3,NSMethod="exhaustive",Distance="minkowski")
specifies a classifier for three-nearest neighbors using the nearest neighbor search
method and the Minkowski metric.
Note
You cannot use any cross-validation name-value argument together with the
OptimizeHyperparameters name-value argument. You can modify the
cross-validation for OptimizeHyperparameters only by using the
HyperparameterOptimizationOptions name-value argument.
Model Parameters
Tie-breaking algorithm used by the predict method if multiple classes have the
same smallest cost, specified as one of the following:
"smallest"— Use the smallest index among tied groups."nearest"— Use the class with the nearest neighbor among tied groups."random"— Use a random tiebreaker among tied groups.
By default, ties occur when multiple classes have the same number of nearest points among the k nearest neighbors.
Example: BreakTies="nearest"
Maximum number of data points in the leaf node of the Kd-tree, specified
as a positive integer value. This argument is meaningful only when
NSMethod is "kdtree".
Example: BucketSize=40
Data Types: single | double
Since R2025a
Size in megabytes of the cache allocated for the Gram matrix,
specified as "maximal" or a positive scalar. The
software can use CacheSize only when the
Distance value begins with
fast.
If the CacheSize value is
"maximal", then during prediction, the software
attempts to allocate enough memory for the Gram matrix, whose size is
n-by-m, where
n is the number of rows in the training predictor
data (X or Tbl), and
m is the number of rows in the test predictor
data. The cache size does not have to be large enough for the Gram
matrix, but must be at least large enough to hold an
n-by-1 vector. Otherwise, the software uses the
regular algorithm for computing the Euclidean distance.
If the Distance value begins with
fast and CacheSize is too
large or is "maximal", then the software might
attempt to allocate a Gram matrix that exceeds the available memory. In
this case, the software issues an error.
Example: CacheSize="maximal"
Data Types: single | double | char | string
Categorical predictor flag, specified as one of the following:
"all"— All predictors are categorical.[]— No predictors are categorical.
The predictor data for fitcknn must be either all continuous
or all categorical.
If the predictor data is in a table (
Tbl),fitcknnassumes that a variable is categorical if it is a logical vector, categorical vector, character array, string array, or cell array of character vectors. IfTblincludes both continuous and categorical values, then you must specify the value ofCategoricalPredictorsso thatfitcknncan determine how to treat all predictors, as either continuous or categorical variables.If the predictor data is a matrix (
X),fitcknnassumes that all predictors are continuous. To identify all predictors inXas categorical, specifyCategoricalPredictorsas"all".
When you set CategoricalPredictors to "all", the
default Distance is "hamming".
Example: CategoricalPredictors="all"
Names of classes to use for training, specified as a categorical, character, or string
array; a logical or numeric vector; or a cell array of character vectors.
ClassNames must have the same data type as the response variable
in Tbl or Y.
If ClassNames is a character array, then each element must correspond to one row of the array.
Use ClassNames to:
Specify the order of the classes during training.
Specify the order of any input or output argument dimension that corresponds to the class order. For example, use
ClassNamesto specify the order of the dimensions ofCostor the column order of classification scores returned bypredict.Select a subset of classes for training. For example, suppose that the set of all distinct class names in
Yis["a","b","c"]. To train the model using observations from classes"a"and"c"only, specifyClassNames=["a","c"].
The default value for ClassNames is the set of all distinct class names in the response variable in Tbl or Y.
Example: ClassNames=["b","g"]
Data Types: categorical | char | string | logical | single | double | cell
Cost of misclassification of a point, specified as one of the following:
Square matrix, where
Cost(i,j)is the cost of classifying a point into classjif its true class isi(that is, the rows correspond to the true class and the columns correspond to the predicted class). To specify the class order for the corresponding rows and columns ofCost, additionally specify theClassNamesname-value pair argument.Structure
Shaving two fields:S.ClassNamescontaining the group names as a variable of the same type asY, andS.ClassificationCostscontaining the cost matrix.
The default is Cost(i,j)=1 if i~=j,
and Cost(i,j)=0 if i=j.
Data Types: single | double | struct
Covariance matrix, specified as a positive definite matrix of scalar values representing the
covariance matrix when computing the Mahalanobis distance. This argument is only valid
when Distance is "mahalanobis".
You cannot simultaneously specify Standardize and either of
Scale or Cov.
Data Types: single | double
Distance metric, specified as a valid distance metric name or function handle. The allowable
distance metric names depend on your choice of a neighbor-searcher method (see
NSMethod).
NSMethod Value | Distance Metric Names |
|---|---|
"exhaustive" | Any distance metric of ExhaustiveSearcher |
"kdtree" | "cityblock", "chebychev",
"euclidean", or "minkowski" |
This table includes valid distance metrics of ExhaustiveSearcher.
| Distance Metric Names | Description |
|---|---|
"cityblock" | City block distance. |
"chebychev" | Chebychev distance (maximum coordinate difference). |
"correlation" | One minus the sample linear correlation between observations (treated as sequences of values). |
"cosine" | One minus the cosine of the included angle between observations (treated as vectors). |
"euclidean" | Euclidean distance. |
| Euclidean distance computed by using an alternative algorithm that saves time
when the number of predictors is at least 10. In some cases, this faster algorithm can
reduce accuracy. Algorithms starting with fast do not support
sparse data. For details, see Fast Euclidean Distance Algorithm. |
| Standardized Euclidean distance computed by using an alternative algorithm that
saves time when the number of predictors is at least 10. In some cases, this faster
algorithm can reduce accuracy. Algorithms starting with fast do not
support sparse data. For details, see Fast Euclidean Distance Algorithm. |
"hamming" | Hamming distance, percentage of coordinates that differ. |
"jaccard" | One minus the Jaccard coefficient, the percentage of nonzero coordinates that differ. |
"mahalanobis" | Mahalanobis distance, computed using a positive definite covariance matrix
C. The default value of C is the sample
covariance matrix of X, as computed by
cov(X,"omitrows"). To specify a different value for
C, use the Cov name-value argument. |
"minkowski" | Minkowski distance. The default exponent is 2. To specify a different
exponent, use the Exponent name-value argument. |
"seuclidean" | Standardized Euclidean distance. Each coordinate difference between X
and a query point is scaled, meaning divided by a scale value S.
The default value of S is the standard deviation computed from
X, S = std(X,"omitnan"). To
specify another value for S, use the Scale
name-value argument. |
"spearman" | One minus the sample Spearman's rank correlation between observations (treated as sequences of values). |
@ |
Distance function handle. function D2 = distfun(ZI,ZJ) % calculation of distance ...
|
If you specify CategoricalPredictors as "all", then
the default distance metric is "hamming". Otherwise, the default distance
metric is "euclidean".
For more information, see Distance Metrics.
Example: Distance="minkowski"
Data Types: char | string | function_handle
Distance weighting function, specified as a function handle or one of the values in this table.
| Value | Description |
|---|---|
"equal" | No weighting |
"inverse" | Weight is 1/distance |
"squaredinverse" | Weight is 1/distance2 |
@ | fcn is a function that accepts a matrix of nonnegative distances,
and returns a matrix the same size containing nonnegative distance
weights. For example, "squaredinverse" is equivalent
to @(d)d.^(-2). |
Example: DistanceWeight="inverse"
Data Types: char | string | function_handle
Minkowski distance exponent, specified as a positive scalar value. This argument is only valid
when Distance is "minkowski".
Example: Exponent=3
Data Types: single | double
Tie inclusion flag, specified as a logical value indicating whether predict includes all the neighbors whose distance values are equal to the
kth smallest distance. If IncludeTies is
true, predict includes all these neighbors.
Otherwise, predict uses exactly k
neighbors.
Example: IncludeTies=true
Data Types: logical
Nearest neighbor search method, specified as "kdtree" or
"exhaustive".
"kdtree"— Creates and uses a Kd-tree to find nearest neighbors."kdtree"is valid when the distance metric is one of the following:"euclidean""cityblock""minkowski""chebychev"
"exhaustive"— Uses the exhaustive search algorithm. When predicting the class of a new pointxnew, the software computes the distance values from all points inXtoxnewto find nearest neighbors.
The default is "kdtree" when X has
10 or fewer columns, X is not sparse or a
gpuArray, and the distance metric is a "kdtree"
type; otherwise, "exhaustive".
Example: NSMethod="exhaustive"
Number of nearest neighbors in X to find for classifying each point when
predicting, specified as a positive integer value.
Example: NumNeighbors=3
Data Types: single | double
Predictor variable names, specified as a string array of unique names or cell array of unique
character vectors. The functionality of PredictorNames depends on the
way you supply the training data.
If you supply
XandY, then you can usePredictorNamesto assign names to the predictor variables inX.The order of the names in
PredictorNamesmust correspond to the column order ofX. That is,PredictorNames{1}is the name ofX(:,1),PredictorNames{2}is the name ofX(:,2), and so on. Also,size(X,2)andnumel(PredictorNames)must be equal.By default,
PredictorNamesis{'x1','x2',...}.
If you supply
Tbl, then you can usePredictorNamesto choose which predictor variables to use in training. That is,fitcknnuses only the predictor variables inPredictorNamesand the response variable during training.PredictorNamesmust be a subset ofTbl.Properties.VariableNamesand cannot include the name of the response variable.By default,
PredictorNamescontains the names of all predictor variables.A good practice is to specify the predictors for training using either
PredictorNamesorformula, but not both.
Example: PredictorNames=["SepalLength","SepalWidth","PetalLength","PetalWidth"]
Data Types: string | cell
Prior probabilities for each class, specified as a value in this table.
| Value | Description |
|---|---|
"empirical" | The class prior probabilities are the class relative frequencies
in Y. |
"uniform" | All class prior probabilities are equal to 1/K, where K is the number of classes. |
| numeric vector | Each element is a class prior probability. Order the elements
according to Mdl.ClassNames or
specify the order using the ClassNames name-value
pair argument. The software normalizes the elements such that they
sum to 1. |
| structure | A structure
|
If you set values for both Weights and Prior, the
weights are renormalized to add up to the value of
the prior probability in the respective
class.
Example: Prior="uniform"
Data Types: char | string | single | double | struct
Response variable name, specified as a character vector or string scalar.
If you supply
Y, then you can useResponseNameto specify a name for the response variable.If you supply
ResponseVarNameorformula, then you cannot useResponseName.
Example: ResponseName="response"
Data Types: char | string
Distance scale, specified as a vector containing nonnegative scalar values with length equal
to the number of columns in X. Each coordinate difference between
X and a query point is scaled by the corresponding element of
Scale. This argument is only valid when
Distance is "seuclidean".
You cannot simultaneously specify Standardize and either of
Scale or Cov.
Data Types: single | double
Score transformation, specified as a character vector, string scalar, or function handle.
This table summarizes the available character vectors and string scalars.
| Value | Description |
|---|---|
"doublelogit" | 1/(1 + e–2x) |
"invlogit" | log(x / (1 – x)) |
"ismax" | Sets the score for the class with the largest score to 1, and sets the scores for all other classes to 0 |
"logit" | 1/(1 + e–x) |
"none" or "identity" | x (no transformation) |
"sign" | –1 for x < 0 0 for x = 0 1 for x > 0 |
"symmetric" | 2x – 1 |
"symmetricismax" | Sets the score for the class with the largest score to 1, and sets the scores for all other classes to –1 |
"symmetriclogit" | 2/(1 + e–x) – 1 |
For a MATLAB function or a function you define, use its function handle for the score transform. The function handle must accept a matrix (the original scores) and return a matrix of the same size (the transformed scores).
Example: ScoreTransform="logit"
Data Types: char | string | function_handle
Flag to standardize the predictors, specified as true
(1) or false
(0).
If you set Standardize=true, then the software centers and scales each
column of the predictor data (X) by the column mean and standard
deviation, respectively.
The software does not standardize categorical predictors, and throws an error if all predictors are categorical.
You cannot simultaneously specify Standardize=true and either of
Scale or Cov.
It is good practice to standardize the predictor data.
Example: Standardize=true
Data Types: logical
Observation weights, specified as a numeric vector of positive values or name of a variable in
Tbl. The software weighs the observations in each row of
X or Tbl with the corresponding value in
Weights. The size of Weights must equal the
number of rows of X or Tbl.
If you specify the input data as a table Tbl, then
Weights can be the name of a variable in Tbl
that contains a numeric vector. In this case, you must specify
Weights as a character vector or string scalar. For example, if
the weights vector W is stored as Tbl.W, then
specify it as "W". Otherwise, the software treats all columns of
Tbl, including W, as predictors or the
response when training the model.
By default, Weights is
ones(, where
n,1)n is the number of observations in X
or Tbl.
The software normalizes Weights to sum up to the value of the prior
probability in the respective class. Inf weights are not supported.
Data Types: double | single | char | string
Cross-Validation Options
Cross-validation flag, specified as "on" or
"off".
If you specify "on", then the software implements 10-fold
cross-validation.
To override this cross-validation setting, use one of these name-value arguments:
CVPartition, Holdout,
KFold, or Leaveout. To create a
cross-validated model, you can use one cross-validation name-value argument at a time
only.
Alternatively, cross validate Mdl later using the crossval
method.
Example: Crossval="on"
Cross-validation partition, specified as a cvpartition object that specifies the type of cross-validation and the
indexing for the training and validation sets.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: Suppose you create a random partition for 5-fold cross-validation on 500
observations by using cvp = cvpartition(500,KFold=5). Then, you can
specify the cross-validation partition by setting
CVPartition=cvp.
Fraction of the data used for holdout validation, specified as a scalar value in the range
(0,1). If you specify Holdout=p, then the software completes these
steps:
Randomly select and reserve
p*100% of the data as validation data, and train the model using the rest of the data.Store the compact trained model in the
Trainedproperty of the cross-validated model.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: Holdout=0.1
Data Types: double | single
Number of folds to use in the cross-validated model, specified as a positive integer value
greater than 1. If you specify KFold=k, then the software completes
these steps:
Randomly partition the data into
ksets.For each set, reserve the set as validation data, and train the model using the other
k– 1 sets.Store the
kcompact trained models in ak-by-1 cell vector in theTrainedproperty of the cross-validated model.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: KFold=5
Data Types: single | double
Leave-one-out cross-validation flag, specified as "on" or
"off". If you specify Leaveout="on", then for
each of the n observations (where n is the number
of observations, excluding missing observations, specified in the
NumObservations property of the model), the software completes
these steps:
Reserve the one observation as validation data, and train the model using the other n – 1 observations.
Store the n compact trained models in an n-by-1 cell vector in the
Trainedproperty of the cross-validated model.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: Leaveout="on"
Data Types: char | string
Hyperparameter Optimization Options
Parameters to optimize, specified as one of the following:
"none"— Do not optimize."auto"— Use["Distance","NumNeighbors","Standardize"]."all"— Optimize all eligible parameters.String array or cell array of eligible parameter names.
Vector of
optimizableVariableobjects, typically the output ofhyperparameters.
The optimization attempts to minimize the cross-validation loss
(error) for fitcknn by varying the parameters. To control the
cross-validation type and other aspects of the optimization, use the
HyperparameterOptimizationOptions name-value argument. When you use
HyperparameterOptimizationOptions, you can use the (compact) model size
instead of the cross-validation loss as the optimization objective by setting the
ConstraintType and ConstraintBounds options.
Note
The values of OptimizeHyperparameters override any values you
specify using other name-value arguments. For example, setting
OptimizeHyperparameters to "auto" causes
fitcknn to optimize hyperparameters corresponding to the
"auto" option and to ignore any specified values for the
hyperparameters.
The eligible parameters for fitcknn are:
Distance—fitcknnsearches among"cityblock","chebychev","correlation","cosine","euclidean","hamming","jaccard","mahalanobis","minkowski","seuclidean", and"spearman".DistanceWeight—fitcknnsearches among"equal","inverse", and"squaredinverse".Exponent—fitcknnsearches among positive real values, by default in the range[0.5,3].NumNeighbors—fitcknnsearches among positive integer values, by default log-scaled in the range[1,max(2,round(NumObservations/2))].Standardize—fitcknnsearches among the values"true"and"false".
Set nondefault parameters by passing a vector of
optimizableVariable objects that have nondefault
values. For example,
load fisheriris params = hyperparameters("fitcknn",meas,species); params(1).Range = [1,20];
Pass params as the value of
OptimizeHyperparameters.
By default, the iterative display appears at the command line,
and plots appear according to the number of hyperparameters in the optimization. For the
optimization and plots, the objective function is the misclassification rate. To control the
iterative display, set the Verbose option of the
HyperparameterOptimizationOptions name-value argument. To control the
plots, set the ShowPlots field of the
HyperparameterOptimizationOptions name-value argument.
For an example, see Optimize KNN Classifier.
Example: OptimizeHyperparameters="auto"
Options for optimization, specified as a HyperparameterOptimizationOptions object or a structure. This argument
modifies the effect of the OptimizeHyperparameters name-value
argument. If you specify HyperparameterOptimizationOptions, you must
also specify OptimizeHyperparameters. All the options are optional.
However, you must set ConstraintBounds and
ConstraintType to return
AggregateOptimizationResults. The options that you can set in a
structure are the same as those in the
HyperparameterOptimizationOptions object.
| Option | Values | Default |
|---|---|---|
Optimizer |
| "bayesopt" |
ConstraintBounds | Constraint bounds for N optimization problems, specified as an N-by-2 numeric matrix or | [] |
ConstraintTarget | Constraint target for the optimization problems, specified as | If you specify ConstraintBounds and ConstraintType, then the default value is "matlab". Otherwise, the default value is []. |
ConstraintType | Constraint type for the optimization problems, specified as | [] |
AcquisitionFunctionName | Type of acquisition function:
Acquisition functions whose names include | "expected-improvement-per-second-plus" |
LossFun | Type of validation loss to optimize, specified as "auto",
"classifcost", "classiferror", or
"mincost". In the case of
fitcknn, the "auto" and
"mincost" options are equivalent, and the software uses
the minimal expected misclassification cost. "classifcost"
indicates to use the observed misclassification cost, and
"classiferror" indicates to use the misclassified rate in
decimal. | "auto" |
MaxObjectiveEvaluations | Maximum number of objective function evaluations. If you specify multiple optimization problems using ConstraintBounds, the value of MaxObjectiveEvaluations applies to each optimization problem individually. | 30 for "bayesopt" and "randomsearch", and the entire grid for "gridsearch" |
MaxTime | Time limit for the optimization, specified as a nonnegative real scalar. The time limit is in seconds, as measured by | Inf |
NumGridDivisions | For Optimizer="gridsearch", the number of values in each dimension. The value can be a vector of positive integers giving the number of values for each dimension, or a scalar that applies to all dimensions. The software ignores this option for categorical variables. | 10 |
ShowPlots | Logical value indicating whether to show plots of the optimization progress. If this option
is true, the software plots the best observed objective
function value against the iteration number. If you use Bayesian optimization
(Optimizer="bayesopt"), the software
also plots the best estimated objective function value. The best observed
objective function values and best estimated objective function values
correspond to the values in the BestSoFar (observed) and
BestSoFar (estim.) columns of the iterative display,
respectively. You can find these values in the properties ObjectiveMinimumTrace and EstimatedObjectiveMinimumTrace of the
SupervisedLearningBayesianOptimization object. If the
problem includes one or two optimization parameters for Bayesian optimization,
then ShowPlots also plots a model of the objective function
against the parameters. | true |
SaveIntermediateResults | Logical value indicating whether to save the optimization results. If this option is
true, the software overwrites a workspace variable named
SupervisedLearningBayesoptResults at each iteration. The
variable is a SupervisedLearningBayesianOptimization object. If you specify
multiple optimization problems using ConstraintBounds, the
workspace variable is an AggregateBayesianOptimization object named
AggregateBayesoptResults. | false |
Verbose | Display level at the command line:
For details, see the | 1 |
UseParallel | Logical value indicating whether to run the Bayesian optimization in parallel, which requires Parallel Computing Toolbox™. Due to the nonreproducibility of parallel timing, parallel Bayesian optimization does not necessarily yield reproducible results. For details, see Parallel Bayesian Optimization. | false |
Repartition | Logical value indicating whether to repartition the cross-validation at every iteration. If this option is A value of | false |
| Specify only one of the following three options. | ||
CVPartition | cvpartition object created by cvpartition | KFold=5 if you do not specify a cross-validation option |
Holdout | Scalar in the range (0,1) representing the holdout fraction | |
KFold | Integer greater than 1 | |
Example: HyperparameterOptimizationOptions=struct(UseParallel=true)
Output Arguments
Trained k-nearest neighbor classification model,
returned as a ClassificationKNN object, a
ClassificationPartitionedModel
object, or a cell array of model objects.
If you set any of the name-value arguments
CrossVal,CVPartition,Holdout,KFold, orLeaveout, thenMdlis aClassificationPartitionedModelobject.If you specify
OptimizeHyperparametersand set theConstraintTypeandConstraintBoundsoptions ofHyperparameterOptimizationOptions, thenMdlis an N-by-1 cell array of model objects, where N is equal to the number of rows inConstraintBounds. If none of the optimization problems yields a feasible model, then each cell array value is[].Otherwise,
Mdlis aClassificationKNNmodel object.
To reference properties of a model object, use dot notation.
Aggregate optimization results for multiple optimization problems, returned as an AggregateBayesianOptimization object. To return
AggregateOptimizationResults, you must specify
OptimizeHyperparameters and
HyperparameterOptimizationOptions. You must also specify the
ConstraintType and ConstraintBounds
options of HyperparameterOptimizationOptions. For an example that
shows how to produce this output, see Hyperparameter Optimization with Multiple Constraint Bounds.
Tips
After training a model, you can generate C/C++ code that predicts labels for new data. Generating C/C++ code requires MATLAB Coder™. For details, see Introduction to Code Generation for Statistics and Machine Learning Functions.
Algorithms
NaNs or <undefined>s indicate missing
observations. The following describes the behavior of fitcknn
when the data set or weights contain missing observations.
If any value of
Yor any weight is missing, thenfitcknnremoves those values fromY, the weights, and the corresponding rows ofXfrom the data. The software renormalizes the weights to sum to1.If you specify to standardize predictors (
Standardize=true) or the standardized Euclidean distance (Distance="seuclidean") without a scale, thenfitcknnremoves missing observations from individual predictors before computing the mean and standard deviation. In other words, the software implementsmeanandstdwith the"omitnan"option on each predictor.If you specify the Mahalanobis distance (
Distance="mahalanobis") without its covariance matrix, thenfitcknnremoves rows ofXthat contain at least one missing value. In other words, the software implementscovwith the"omitrows"option on the predictor matrixX.
If you specify the
Cost,Prior, andWeightsname-value arguments, the output model object stores the specified values in theCost,Prior, andWproperties, respectively. TheCostproperty stores the user-specified cost matrix as is. ThePriorandWproperties store the prior probabilities and observation weights, respectively, after normalization. For details, see Misclassification Cost Matrix, Prior Probabilities, and Observation Weights.The software uses the
Costproperty for prediction, but not training. Therefore,Costis not read-only; you can change the property value by using dot notation after creating the trained model.Suppose that you set
Standardize=true.If you also specify the
PriororWeightsname-value argument, thenfitcknnstandardizes the predictors using their corresponding weighted means and weighted standard deviations. Specifically,fitcknnstandardizes the predictor j using-
xjk is observation k (row) of predictor j (column).
-
If you also set
Distance="mahalanobis"orDistance="seuclidean", then you cannot specifyScaleorCov. Instead, the software:Computes the means and standard deviations of each predictor.
Standardizes the data using the results of step 1.
Computes the distance parameter values using their respective default.
If you specify
Scaleand either ofPriororWeights, then the software scales observed distances by the weighted standard deviations.If you specify
Covand either ofPriororWeights, then the software applies the weighted covariance matrix to the distances. In other words,
The Distance argument values that begin with fast
(such as "fasteuclidean" and "fastseuclidean")
calculate Euclidean distances using an algorithm that uses extra memory to save
computational time. This algorithm is named "Euclidean Distance Matrix Trick" in Albanie
[1] and elsewhere. Internal
testing shows that this algorithm saves time when the number of predictors is at least 10.
Algorithms starting with fast do not support sparse data.
To find the matrix D of distances between all the points xi and xj, where each xi has n variables, the algorithm computes distance using the final line in the following equations:
The matrix in the last line of the equations is called the Gram matrix. Computing the set of squared distances is faster, but slightly less numerically stable, when you compute and use the Gram matrix instead of computing the squared distances by squaring and summing. For more details, see Albanie [1].
To store the Gram matrix, the software uses a cache with the default size of
1e3 megabytes. You can set the cache size using the
CacheSize name-value argument. If the value of
CacheSize is too large or "maximal", then the
software might try to allocate a Gram matrix that exceeds the available memory. In this
case, the software issues an error.
ClassificationKNN predicts the
classification of a point xnew using a procedure equivalent to
this:
Find the
NumNeighborspoints in the training setXthat are nearest toxnew.Find the
NumNeighborsresponse valuesYto those nearest points.Assign the classification label
ynewthat has the largest posterior probability among the values inY.
For details, see Posterior Probability in the predict documentation.
Alternatives
Although fitcknn can train a multiclass KNN classifier, you can
reduce a multiclass learning problem to a series of KNN binary learners using fitcecoc.
References
[1] Albanie, Samuel. Euclidean Distance Matrix Trick. June, 2019. Available at https://samuelalbanie.com/files/Euclidean_distance_trick.pdf.
Extended Capabilities
To perform parallel hyperparameter optimization, use the UseParallel=true
option in the HyperparameterOptimizationOptions name-value argument in
the call to the fitcknn function.
For more information on parallel hyperparameter optimization, see Parallel Bayesian Optimization.
For general information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
Usage notes and limitations:
By default,
fitcknnuses the exhaustive nearest neighbor search algorithm forgpuArrayinput arguments.You cannot specify the name-value argument
NSMethodas"kdtree".You cannot specify the name-value argument
Distanceas"fasteuclidean","fastseuclidean", or a function handle.You cannot specify the name-value argument
IncludeTiesastrue.fitcknnfits the model on a GPU if one of the following applies:The input argument
Xis agpuArrayobject.The input argument
TblcontainsgpuArraypredictor variables.
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2014aYou can optimize classification model hyperparameters with respect to misclassification
cost. When you use a classification fit function, specify the
OptimizeHyperparameters and
HyperparameterOptimizationOptions name-value arguments. In the
HyperparameterOptimizationOptions structure or object, set the
LossFun value to "classifcost",
"mincost", or "auto-cost", depending on the
classification fit function.
Depending on the type of supervised learning fit function you use to perform
hyperparameter optimization, you can set the LossFun value to
"auto-cost", "classifcost",
"classiferror", "mincost",
"mse", or "quantile".
fitcknn defaults to serial hyperparameter optimization when
HyperparameterOptimizationOptions includes
UseParallel=true and the software cannot open a parallel pool.
In previous releases, the software issues an error under these circumstances.
Starting in R2023b, when you specify "auto" as the OptimizeHyperparameters value, fitcknn includes Standardize as an optimizable hyperparameter.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)