Introduction to Fairness in Binary Classification
The functions fairnessMetrics, fairnessWeights,
disparateImpactRemover, and fairnessThresholder in
Statistics and Machine Learning Toolbox™ allow you to detect and mitigate societal bias in binary classification. First,
use fairnessMetrics to evaluate the fairness of a data set or
classification model with bias and group metrics. Then, use fairnessWeights to
reweight observations, disparateImpactRemover to remove the disparate impact of a sensitive attribute,
or fairnessThresholder
to optimize the classification threshold.
fairnessMetrics— ThefairnessMetricsfunction computes fairness metrics (bias metrics and group metrics) for a data set or binary classification model with respect to sensitive attributes. The data-level evaluation examines binary, true labels of the data. The model-level evaluation examines the predicted labels returned by one or more binary classification models, using both true labels and predicted labels. You can use the metrics to determine if your data or models contain bias toward a group within each sensitive attribute.fairnessWeights— ThefairnessWeightsfunction computes fairness weights with respect to a sensitive attribute and the response variable. For every combination of a group in the sensitive attribute and a class label in the response variable, the software computes a weight value. The function then assigns each observation its corresponding weight. The returned weights introduce fairness across the sensitive attribute groups. Pass the weights to an appropriate training function, such asfitcsvm, using theWeightsname-value argument.disparateImpactRemover— ThedisparateImpactRemoverfunction tries to remove the disparate impact of a sensitive attribute on model predictions by using the sensitive attribute to transform the continuous predictors in the data set. The function returns the transformed data set and adisparateImpactRemoverobject that contains the transformation. Pass the transformed data set to an appropriate training function, such asfitcsvm, and pass the object to thetransformobject function to apply the transformation to a new data set, such as a test data set.fairnessThresholder— ThefairnessThresholderfunction searches for an optimal score threshold to maximize accuracy while satisfying fairness bounds. For observations in the critical region below the optimal threshold, the function adjusts the labels so that the fairness constraints hold for the reference and nonreference groups in the sensitive attribute. After you create afairnessThresholderobject, you can use thepredictandlossobject functions on new data to predict fairness labels and calculate the classification loss, respectively.
Reduce Statistical Parity Difference Using Fairness Weights
Train a neural network model, and compute the statistical parity difference (SPD) for each group in the sensitive attribute. To reduce the SPD values, compute fairness weights, and retrain the neural network model.
Read the sample file CreditRating_Historical.dat into a table. The predictor data consists of financial ratios and industry sector information for a list of corporate customers. The response variable consists of credit ratings assigned by a rating agency.
creditrating = readtable("CreditRating_Historical.dat");Because each value in the ID variable is a unique customer ID—that is, length(unique(creditrating.ID)) is equal to the number of observations in creditrating—the ID variable is a poor predictor. Remove the ID variable from the table, and convert the Industry variable to a categorical variable.
creditrating.ID = []; creditrating.Industry = categorical(creditrating.Industry);
In the Rating response variable, combine the AAA, AA, A, and BBB ratings into a category of "good" ratings, and the BB, B, and CCC ratings into a category of "poor" ratings.
Rating = categorical(creditrating.Rating); Rating = mergecats(Rating,["AAA","AA","A","BBB"],"good"); Rating = mergecats(Rating,["BB","B","CCC"],"poor"); creditrating.Rating = Rating;
Train a neural network model on the creditrating data. For better results, standardize the predictors before fitting the model. Use the trained model to predict labels for the training data set.
rng("default") % For reproducibility netMdl = fitcnet(creditrating,"Rating",Standardize=true); netPredictions = predict(netMdl,creditrating);
Compute fairness metrics with respect to the Industry sensitive attribute by using the model predictions. In particular, find the statistical parity difference (SPD) for each group in Industry.
netMetricsResults = fairnessMetrics(creditrating,"Rating", ... SensitiveAttributeNames="Industry",Predictions=netPredictions); report(netMetricsResults,BiasMetrics="StatisticalParityDifference")
ans=12×4 table
ModelNames SensitiveAttributeNames Groups StatisticalParityDifference
__________ _______________________ ______ ___________________________
Model1 Industry 1 0.065724
Model1 Industry 2 0.078449
Model1 Industry 3 0
Model1 Industry 4 0.097937
Model1 Industry 5 0.05251
Model1 Industry 6 -0.0061672
Model1 Industry 7 0.030037
Model1 Industry 8 0.087863
Model1 Industry 9 0.16367
Model1 Industry 10 0.15076
Model1 Industry 11 0.019402
Model1 Industry 12 0.032187
To better understand the distribution of SPD values, plot the values using a box plot.
spdValues = netMetricsResults.BiasMetrics.StatisticalParityDifference; boxchart(spdValues) ylabel("Statistical Parity Difference") title("Distribution of Statistical Parity Differences")
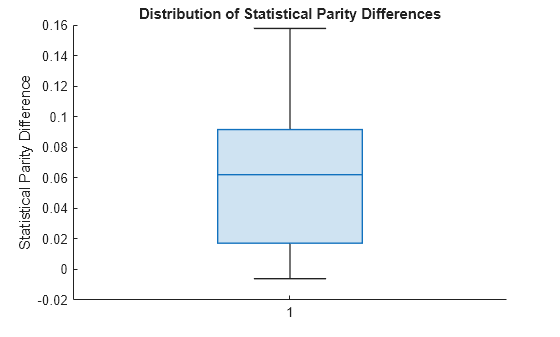
The median SPD value is around 0.06, which is higher than the value 0 of a fair model.
Compute fairness weights, and refit a neural network model using the weights. As before, standardize the predictors. Then, predict labels for the training data by using the new model.
weights = fairnessWeights(creditrating,"Industry","Rating"); rng("default") % For reproducibility newNetMdl = fitcnet(creditrating,"Rating",Weights=weights, ... Standardize=true); newNetPredictions = predict(newNetMdl,creditrating);
Compute the new SPD values.
newNetMetricsResults = fairnessMetrics(creditrating,"Rating", ... SensitiveAttributeNames="Industry",Predictions=newNetPredictions); report(newNetMetricsResults,BiasMetrics="StatisticalParityDifference")
ans=12×4 table
ModelNames SensitiveAttributeNames Groups StatisticalParityDifference
__________ _______________________ ______ ___________________________
Model1 Industry 1 0.051528
Model1 Industry 2 0.070411
Model1 Industry 3 0
Model1 Industry 4 0.06844
Model1 Industry 5 0.029635
Model1 Industry 6 -0.0051521
Model1 Industry 7 0.016443
Model1 Industry 8 0.070997
Model1 Industry 9 0.11324
Model1 Industry 10 0.11538
Model1 Industry 11 0.026125
Model1 Industry 12 0.0090789
Display the two distributions of SPD values. The left box plot shows the SPD values computed using the original model. The right box plot shows the SPD values computed using the new model trained with fairness weights.
spdValuesUpdated = newNetMetricsResults.BiasMetrics.StatisticalParityDifference; boxchart([spdValues spdValuesUpdated]) xticklabels(["Without Weights","With Weights"]) ylabel("Statistical Parity Difference") title("Distribution of Statistical Parity Differences")
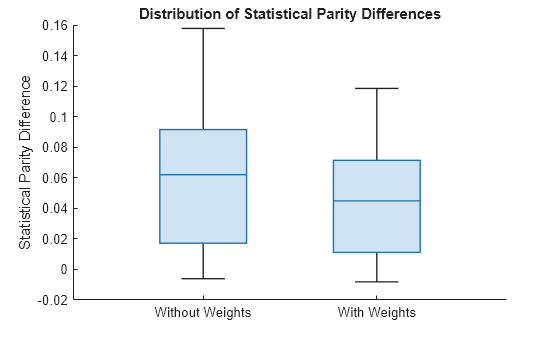
The new SPD values have a median around 0.04, which is closer to 0 than the previous median of 0.06. The maximum value of the new SPD values, which is around 0.11, is also closer to 0 than the previous maximum value, which is around 0.16.
Reduce Disparate Impact of Predictions
Train a binary classifier, classify test data using the model, and compute the disparate impact for each group in the sensitive attribute. To reduce the disparate impact values, use disparateImpactRemover, and then retrain the binary classifier. Transform the test data set, reclassify the observations, and compute the disparate impact values.
Load the sample data census1994, which contains the training data adultdata and the test data adulttest. The data sets consist of demographic information from the US Census Bureau that can be used to predict whether an individual makes over $50,000 per year. Preview the first few rows of the training data set.
load census1994
head(adultdata) age workClass fnlwgt education education_num marital_status occupation relationship race sex capital_gain capital_loss hours_per_week native_country salary
___ ________________ __________ _________ _____________ _____________________ _________________ _____________ _____ ______ ____________ ____________ ______________ ______________ ______
39 State-gov 77516 Bachelors 13 Never-married Adm-clerical Not-in-family White Male 2174 0 40 United-States <=50K
50 Self-emp-not-inc 83311 Bachelors 13 Married-civ-spouse Exec-managerial Husband White Male 0 0 13 United-States <=50K
38 Private 2.1565e+05 HS-grad 9 Divorced Handlers-cleaners Not-in-family White Male 0 0 40 United-States <=50K
53 Private 2.3472e+05 11th 7 Married-civ-spouse Handlers-cleaners Husband Black Male 0 0 40 United-States <=50K
28 Private 3.3841e+05 Bachelors 13 Married-civ-spouse Prof-specialty Wife Black Female 0 0 40 Cuba <=50K
37 Private 2.8458e+05 Masters 14 Married-civ-spouse Exec-managerial Wife White Female 0 0 40 United-States <=50K
49 Private 1.6019e+05 9th 5 Married-spouse-absent Other-service Not-in-family Black Female 0 0 16 Jamaica <=50K
52 Self-emp-not-inc 2.0964e+05 HS-grad 9 Married-civ-spouse Exec-managerial Husband White Male 0 0 45 United-States >50K
Each row contains the demographic information for one adult. The last column salary shows whether a person has a salary less than or equal to $50,000 per year or greater than $50,000 per year.
Remove observations from adultdata and adulttest that contain missing values.
adultdata = rmmissing(adultdata); adulttest = rmmissing(adulttest);
Specify the continuous numeric predictors to use for model training.
predictors = ["age","education_num","capital_gain","capital_loss", ... "hours_per_week"];
Train an ensemble classifier using the training set adultdata. Specify salary as the response variable and fnlwgt as the observation weights. Because the training set is imbalanced, use the RUSBoost algorithm. After training the model, predict the salary (class label) of the observations in the test set adulttest.
rng("default") % For reproducibility mdl = fitcensemble(adultdata,"salary",Weights="fnlwgt", ... PredictorNames=predictors,Method="RUSBoost"); labels = predict(mdl,adulttest);
Transform the training set predictors by using the race sensitive attribute.
[remover,newadultdata] = disparateImpactRemover(adultdata, ... "race",PredictorNames=predictors); remover
remover =
disparateImpactRemover with properties:
RepairFraction: 1
PredictorNames: {'age' 'education_num' 'capital_gain' 'capital_loss' 'hours_per_week'}
SensitiveAttribute: 'race'
remover is a disparateImpactRemover object, which contains the transformation of the remover.PredictorNames predictors with respect to the remover.SensitiveAttribute variable.
Apply the same transformation stored in remover to the test set predictors. Note: You must transform both the training and test data sets before passing them to a classifier.
newadulttest = transform(remover,adulttest, ...
PredictorNames=predictors);Train the same type of ensemble classifier as mdl, but use the transformed predictor data. As before, predict the salary (class label) of the observations in the test set adulttest.
rng("default") % For reproducibility newMdl = fitcensemble(newadultdata,"salary",Weights="fnlwgt", ... PredictorNames=predictors,Method="RUSBoost"); newLabels = predict(newMdl,newadulttest);
Compare the disparate impact values for the predictions made by the original model (mdl) and the predictions made by the model trained with the transformed data (newMdl). For each group in the sensitive attribute, the disparate impact value is the proportion of predictions in that group with a positive class value () divided by the proportion of predictions in the reference group with a positive class value (). An ideal classifier makes predictions where, for each group, is close to (that is, where the disparate impact value is close to 1).
Compute the disparate impact values for the mdl predictions and the newMdl predictions by using fairnessMetrics. Include the observation weights. You can use the report object function to display bias metrics, such as disparate impact, that are stored in the metricsResults object.
metricsResults = fairnessMetrics(adulttest,"salary", ... SensitiveAttributeNames="race",Predictions=[labels,newLabels], ... Weights="fnlwgt",ModelNames=["Original Model","New Model"]); metricsResults.PositiveClass
ans = categorical
>50K
metricsResults.ReferenceGroup
ans = 'White'
report(metricsResults,BiasMetrics="DisparateImpact")ans=5×5 table
Metrics SensitiveAttributeNames Groups Original Model New Model
_______________ _______________________ __________________ ______________ _________
DisparateImpact race Amer-Indian-Eskimo 0.41702 0.92804
DisparateImpact race Asian-Pac-Islander 1.719 0.9697
DisparateImpact race Black 0.60571 0.66629
DisparateImpact race Other 0.66958 0.86039
DisparateImpact race White 1 1
For the mdl predictions, several of the disparate impact values are below the industry standard of 0.8, and one value is above 1.25. These values indicate bias in the predictions with respect to the positive class >50K and the sensitive attribute race.
The disparate impact values for the newMdl predictions are closer to 1 than the disparate impact values for the mdl predictions. One value is still below 0.8.
Visually compare the disparate impact values by using the bar graph returned by the plot object function.
plot(metricsResults,"DisparateImpact")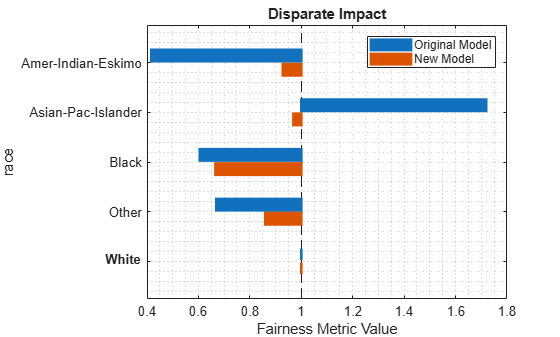
The disparateImpactRemover function seems to have improved the model predictions on the test set with respect to the disparate impact metric.
Check whether the transformed predictors negatively affect the accuracy of the model predictions. Compute the accuracy of the test set predictions for the two models mdl and newMdl.
accuracy = 1-loss(mdl,adulttest,"salary")accuracy = 0.8024
newAccuracy = 1-loss(newMdl,newadulttest,"salary")newAccuracy = 0.7955
The model trained using the transformed predictors (newMdl) achieves similar test set accuracy compared to the model trained with the original predictors (mdl).
See Also
fairnessMetrics | fairnessWeights | disparateImpactRemover | transform | fairnessThresholder | loss | predict