testcholdout
Compare predictive accuracies of two classification models
Syntax
Description
testcholdout statistically assesses
the accuracies of two classification models. The function first compares
their predicted labels against the true labels, and then it detects
whether the difference between the misclassification rates is statistically
significant.
You can assess whether the accuracies of the classification
models are different, or whether one classification model performs
better than another. testcholdout can conduct several McNemar test variations,
including the asymptotic test, the exact-conditional test, and the
mid-p-value test. For cost-sensitive assessment, available
tests include a chi-square test (requires an Optimization Toolbox™ license)
and a likelihood ratio test.
h = testcholdout(YHat1,YHat2,Y)YHat1 and YHat2 have
equal accuracy for predicting the true class labels Y.
The alternative hypothesis is that the labels have unequal accuracy.
h = 1 indicates to reject
the null hypothesis at the 5% significance level. h = 0 indicates
to not reject the null hypothesis at 5% level.
h = testcholdout(YHat1,YHat2,Y,Name,Value)Name,Value pair arguments. For example,
you can specify the type of alternative hypothesis, specify the type
of test, or supply a cost matrix.
Examples
Train two classification models using different algorithms. Conduct a statistical test comparing the misclassification rates of the two models on a held-out set.
Load the ionosphere data set.
load ionosphereCreate a partition that evenly splits the data into training and testing sets.
rng(1); % For reproducibility CVP = cvpartition(Y,'holdout',0.5); idxTrain = training(CVP); % Training-set indices idxTest = test(CVP); % Test-set indices
CVP is a cross-validation partition object that specifies the training and test sets.
Train an SVM model and an ensemble of 100 bagged classification trees. For the SVM model, specify to use the radial basis function kernel and a heuristic procedure to determine the kernel scale.
MdlSVM = fitcsvm(X(idxTrain,:),Y(idxTrain),'Standardize',true,... 'KernelFunction','RBF','KernelScale','auto'); t = templateTree('Reproducible',true); % For reproducibility of random predictor selections MdlBag = fitcensemble(X(idxTrain,:),Y(idxTrain),'Method','Bag','Learners',t);
MdlSVM is a trained ClassificationSVM model. MdlBag is a trained ClassificationBaggedEnsemble model.
Label the test-set observations using the trained models.
YhatSVM = predict(MdlSVM,X(idxTest,:)); YhatBag = predict(MdlBag,X(idxTest,:));
YhatSVM and YhatBag are vectors continuing the predicted class labels of the respective models.
Test whether the two models have equal predictive accuracies.
h = testcholdout(YhatSVM,YhatBag,Y(idxTest))
h = logical
0
h = 0 indicates to not reject the null hypothesis that the two models have equal predictive accuracies.
Train two classification models using the same algorithm, but adjust a hyperparameter to make the algorithm more complex. Conduct a statistical test to assess whether the simpler model has better accuracy in held-out data than the more complex model.
Load the ionosphere data set.
load ionosphere;Create a partition that evenly splits the data into training and testing sets.
rng(1); % For reproducibility CVP = cvpartition(Y,'holdout',0.5); idxTrain = training(CVP); % Training-set indices idxTest = test(CVP); % Test-set indices
CVP is a cross-validation partition object that specifies the training and test sets.
Train two SVM models: one that uses a linear kernel (the default for binary classification) and one that uses the radial basis function kernel. Use the default kernel scale of 1.
MdlLinear = fitcsvm(X(idxTrain,:),Y(idxTrain),'Standardize',true); MdlRBF = fitcsvm(X(idxTrain,:),Y(idxTrain),'Standardize',true,... 'KernelFunction','RBF');
MdlLinear and MdlRBF are trained ClassificationSVM models.
Label the test-set observations using the trained models.
YhatLinear = predict(MdlLinear,X(idxTest,:)); YhatRBF = predict(MdlRBF,X(idxTest,:));
YhatLinear and YhatRBF are vectors continuing the predicted class labels of the respective models.
Test the null hypothesis that the simpler model (MdlLinear) is at most as accurate as the more complex model (MdlRBF). Because the test-set size is large, conduct the asymptotic McNemar test, and compare the results with the mid- p-value test (the cost-insensitive testing default). Request to return p-values and misclassification rates.
Asymp = zeros(4,1); % Preallocation MidP = zeros(4,1); [Asymp(1),Asymp(2),Asymp(3),Asymp(4)] = testcholdout(YhatLinear,YhatRBF,Y(idxTest),... 'Alternative','greater','Test','asymptotic'); [MidP(1),MidP(2),MidP(3),MidP(4)] = testcholdout(YhatLinear,YhatRBF,Y(idxTest),... 'Alternative','greater'); table(Asymp,MidP,'RowNames',{'h' 'p' 'e1' 'e2'})
ans=4×2 table
Asymp MidP
__________ __________
h 1 1
p 7.2801e-09 2.7649e-10
e1 0.13714 0.13714
e2 0.33143 0.33143
The p-value is close to zero for both tests, which indicates strong evidence to reject the null hypothesis that the simpler model is less accurate than the more complex model. No matter what test you specify, testcholdout returns the same type of misclassification measure for both models.
For data sets with imbalanced class representations, or if the false-positive and false-negative costs are imbalanced, you can statistically compare the predictive performance of two classification models by including a cost matrix in the analysis.
Load the arrhythmia data set. Determine the class representations in the data.
load arrhythmia;
Y = categorical(Y);
tabulate(Y); Value Count Percent
1 245 54.20%
2 44 9.73%
3 15 3.32%
4 15 3.32%
5 13 2.88%
6 25 5.53%
7 3 0.66%
8 2 0.44%
9 9 1.99%
10 50 11.06%
14 4 0.88%
15 5 1.11%
16 22 4.87%
There are 16 classes, however some are not represented in the data set (for example, class 13). Most observations are classified as not having arrhythmia (class 1). The data set is highly discrete with imbalanced classes.
Combine all observations with arrhythmia (classes 2 through 15) into one class. Remove those observations with unknown arrhythmia status (class 16) from the data set.
idx = (Y ~= '16'); Y = Y(idx); X = X(idx,:); Y(Y ~= '1') = 'WithArrhythmia'; Y(Y == '1') = 'NoArrhythmia'; Y = removecats(Y);
Create a partition that evenly splits the data into training and test sets.
rng(1); % For reproducibility CVP = cvpartition(Y,'holdout',0.5); idxTrain = training(CVP); % Training-set indices idxTest = test(CVP); % Test-set indices
CVP is a cross-validation partition object that specifies the training and test sets.
Create a cost matrix such that misclassifying a patient with arrhythmia into the "no arrhythmia" class is five times worse than misclassifying a patient without arrhythmia into the arrhythmia class. Classifying correctly incurs no cost. The rows indicate the true class and the columns indicate predicted class. When you conduct a cost-sensitive analysis, a good practice is to specify the order of the classes.
Cost = [0 1;5 0];
ClassNames = {'NoArrhythmia','WithArrhythmia'};Train two boosting ensembles of 50 classification trees, one that uses AdaBoostM1 and another that uses LogitBoost. Because there are missing values in the data set, specify to use surrogate splits. Train the models using the cost matrix.
t = templateTree('Surrogate','on'); numTrees = 50; MdlAda = fitcensemble(X(idxTrain,:),Y(idxTrain),'Method','AdaBoostM1',... 'NumLearningCycles',numTrees,'Learners',t,... 'Cost',Cost,'ClassNames',ClassNames); MdlLogit = fitcensemble(X(idxTrain,:),Y(idxTrain),'Method','LogitBoost',... 'NumLearningCycles',numTrees,'Learners',t,... 'Cost',Cost,'ClassNames',ClassNames);
MdlAda and MdlLogit are trained ClassificationEnsemble models.
Label the test-set observations using the trained models.
YhatAda = predict(MdlAda,X(idxTest,:)); YhatLogit = predict(MdlLogit,X(idxTest,:));
YhatLinear and YhatRBF are vectors containing the predicted class labels of the respective models.
Test whether the AdaBoostM1 ensemble (MdlAda) and the LogitBoost ensemble (MdlLogit) have equal predictive accuracy. Supply the cost matrix. Conduct the asymptotic, likelihood ratio, cost-sensitive test (the default when you pass in a cost matrix). Request to return p-values and misclassification costs.
[h,p,e1,e2] = testcholdout(YhatAda,YhatLogit,Y(idxTest), ... 'Cost',Cost,'ClassNames',ClassNames)
h = logical
0
p = 0.1180
e1 = 0.6698
e2 = 0.8093
h = 0 indicates to not reject the null hypothesis that the two models have equal predictive accuracies.
Input Arguments
Predicted class labels of the first classification model, specified as a categorical, character, or string array, logical or numeric vector, or cell array of character vectors.
If YHat1 is a character array, then each
element must correspond to one row of the array.
YHat1, YHat2, and Y must
have equal lengths.
It is a best practice for YHat1, YHat2,
and Y to share the same data type.
Data Types: categorical | char | string | logical | single | double | cell
Predicted class labels of the second classification model, specified as a categorical, character, or string array, logical or numeric vector, or cell array of character vectors.
If YHat2 is a character array, then each
element must correspond to one row of the array.
YHat1, YHat2, and Y must
have equal lengths.
It is a best practice for YHat1, YHat2,
and Y to share the same data type.
Data Types: categorical | char | string | logical | single | double | cell
True class labels, specified as a categorical, character, or string array, logical or numeric vector, or cell array of character vectors.
If Y is a character array, then each element
must correspond to one row of the array.
YHat1, YHat2, and Y must
have equal lengths.
It is a best practice for YHat1, YHat2,
and Y to share the same data type.
Data Types: categorical | char | string | logical | single | double | cell
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: 'Alternative','greater','Test','asymptotic','Cost',[0
2;1 0] specifies to test whether the first set of first
predicted class labels is more accurate than the second set, to conduct
the asymptotic McNemar test, and to penalize misclassifying observations
with the true label ClassNames{1} twice as much
as for misclassifying observations with the true label ClassNames{2}.
Hypothesis test significance level, specified as the comma-separated
pair consisting of 'Alpha' and a scalar value in
the interval (0,1).
Example: 'Alpha',0.1
Data Types: single | double
Alternative hypothesis to assess, specified as the comma-separated
pair consisting of 'Alternative' and one of the
values listed in the table.
| Value | Alternative hypothesis |
|---|---|
'unequal' (default) | For predicting Y, YHat1 and YHat2 have
unequal accuracies. |
'greater' | For predicting Y, YHat1 is
more accurate than YHat2. |
'less' | For predicting Y, YHat1 is
less accurate than YHat2. |
Example: 'Alternative','greater'
Class names, specified as the comma-separated pair consisting of
'ClassNames' and a categorical, character, or
string array, logical or numeric vector, or cell array of character
vectors. You must set ClassNames using the data
type of Y.
If ClassNames is a character array, then each
element must correspond to one row of the array.
Use ClassNames to:
Specify the order of any input argument dimension that corresponds to class order. For example, use
ClassNamesto specify the order of the dimensions ofCost.Select a subset of classes for testing. For example, suppose that the set of all distinct class names in
Yis{'a','b','c'}. To train and test models using observations from classes'a'and'c'only, specify'ClassNames',{'a','c'}.
The default is the set of all distinct class names in
Y.
Example: 'ClassNames',{'b','g'}
Data Types: single | double | logical | char | string | cell | categorical
Misclassification cost, specified as the comma-separated pair consisting of
'Cost' and a square matrix or structure array.
If you specify the square matrix
Cost, thenCost(i,j)is the cost of classifying a point into classjif its true class isi. That is, the rows correspond to the true class and the columns correspond to the predicted class. To specify the class order for the corresponding rows and columns ofCost, additionally specify theClassNamesname-value pair argument.If you specify the structure
S, thenSmust have two fields:S.ClassNames, which contains the class names as a variable of the same data type asY. You can use this field to specify the order of the classes.S.ClassificationCosts, which contains the cost matrix, with rows and columns ordered as inS.ClassNames.
If you specify Cost, then testcholdout cannot conduct
one-sided, exact, or mid-p tests. You must also specify
'Alternative','unequal','Test','asymptotic'. For cost-sensitive
testing options, see the CostTest name-value pair argument.
A best practice is to supply the same cost matrix used to train the classification models.
The default is Cost(i,j) = 1 if i
~= j, and Cost(i,j) = 0 if i
= j.
Example: 'Cost',[0 1 2 ; 1 0 2; 2 2 0]
Data Types: single | double | struct
Cost-sensitive test type, specified as the comma-separated pair
consisting of 'CostTest' and 'chisquare' or 'likelihood'.
Unless you specify a cost matrix using the Cost name-value
pair argument, testcholdout ignores CostTest.
This table summarizes the available options for cost-sensitive testing.
| Value | Asymptotic test type | Requirements |
|---|---|---|
'chisquare' | Chi-square test | Optimization Toolbox license to implement quadprog (Optimization Toolbox) |
'likelihood' | Likelihood ratio test | None |
For more details, see Cost-Sensitive Testing.
Example: 'CostTest','chisquare'
Test to conduct, specified as the comma-separated pair consisting
of 'Test' and 'asymptotic', 'exact',
and 'midp'. This table summarizes the available
options for cost-insensitive testing.
| Value | Description |
|---|---|
'asymptotic' | Asymptotic McNemar test |
'exact' | Exact-conditional McNemar test |
'midp' (default) | Mid-p-value McNemar test |
For more details, see McNemar Tests.
For cost-sensitive testing, Test must be 'asymptotic'.
When you specify the Cost name-value pair argument,
and choose a cost-sensitive test using the CostTest name-value
pair argument, 'asymptotic' is the default.
Example: 'Test','asymptotic'
Note
NaNs, <undefined> values, empty character vectors
(''), empty strings (""), and
<missing> values indicate missing data values.
testcholdout:
Treats missing values in
YHat1andYHat2as misclassified observations.Removes missing values in
Yand the corresponding values ofYHat1andYHat2
Output Arguments
Hypothesis test result, returned as a logical value.
h = 1 indicates the rejection of the null
hypothesis at the Alpha significance level.
h = 0 indicates failure to reject the null hypothesis at the
Alpha significance level.
Data Types: logical
p-value of the test, returned as a scalar
in the interval [0,1]. p is the probability that
a random test statistic is at least as extreme as the observed test
statistic, given that the null hypothesis is true.
testcholdout estimates p using
the distribution of the test statistic, which varies with the type
of test. For details on test statistics derived from the available
variants of the McNemar test, see McNemar Tests. For details on test statistics derived from
cost-sensitive tests, see Cost-Sensitive Testing.
Classification
loss that summarizes the accuracy of the first set of class
labels (YHat1) predicting the true class labels
(Y), returned as a scalar.
For cost-insensitive testing, e1 is the misclassification
rate. That is, e1 is the proportion of misclassified
observations, which is a scalar in the interval [0,1].
For cost-sensitive testing, e1 is the misclassification
cost. That is, e1 is the weighted average of the
misclassification costs, in which the weights are the respective estimated
proportions of misclassified observations.
Classification
loss that summarizes the accuracy of the second set of class
labels (YHat2) predicting the true class labels
(Y), returned as a scalar.
For cost-insensitive testing, e2 is the misclassification
rate. That is, e2 is the proportion of misclassified
observations, which is a scalar in the interval [0,1].
For cost-sensitive testing, e2 is the misclassification
cost. That is, e2 is the weighted average of the
costs of misclassification, in which the weights are the respective
estimated proportions of misclassified observations.
More About
Conduct cost-sensitive testing when the cost of misclassification is imbalanced. By conducting a cost-sensitive analysis, you can account for the cost imbalance when you train the classification models and when you statistically compare them.
If the cost of misclassification is imbalanced, then the misclassification rate tends to be a poorly performing classification loss. Use misclassification cost instead to compare classification models.
Misclassification costs are often imbalanced in applications. For example, consider classifying subjects based on a set of predictors into two categories: healthy and sick. Misclassifying a sick subject as healthy poses a danger to the subject's life. However, misclassifying a healthy subject as sick typically causes some inconvenience, but does not pose significant danger. In this situation, you assign misclassification costs such that misclassifying a sick subject as healthy is more costly than misclassifying a healthy subject as sick.
The definitions that follow summarize the cost-sensitive tests. In the definitions:
nijk and are the number and estimated proportion of test-sample observations with the following characteristics. k is the true class, i is the label assigned by the first classification model, and j is the label assigned by the second classification model. The unknown true value of is πijk. The test-set sample size is Additionally,
cij is the relative cost of assigning label j to an observation with true class i. cii = 0, cij ≥ 0, and, for at least one (i,j) pair, cij > 0.
All subscripts take on integer values from 1 through K, which is the number of classes.
The expected difference in the misclassification costs of the two classification models is
The hypothesis test is
The available cost-sensitive tests are appropriate for two-tailed testing.
Available asymptotic tests that address imbalanced costs are a chi-square test and a likelihood ratio test.
Chi-square test — The chi-square test statistic is based on the Pearson and Neyman chi-square test statistics, but with a Laplace correction factor to account for any nijk = 0. The test statistic is
If , then reject H0.
are estimated by minimizing under the constraint that δ = 0.
is the χ2 cdf with one degree of freedom evaluated at x.
Likelihood ratio test — The likelihood ratio test is based on Nijk, which are binomial random variables with sample size ntest and success probability πijk. The random variables represent the random number of observations with: true class k, label i assigned by the first classification model, and label j assigned by the second classification model. Jointly, the distribution of the random variables is multinomial.
The test statistic is
If then reject H0.
is the unrestricted MLE of πijk.
is the MLE under the null hypothesis that δ = 0. λ is the solution to
is the χ2 cdf with one degree of freedom evaluated at x.
McNemar Tests are hypothesis tests that compare two population proportions while addressing the issues resulting from two dependent, matched-pair samples.
One way to compare the predictive accuracies of two classification models is:
Partition the data into training and test sets.
Train both classification models using the training set.
Predict class labels using the test set.
Summarize the results in a two-by-two table similar to this figure.
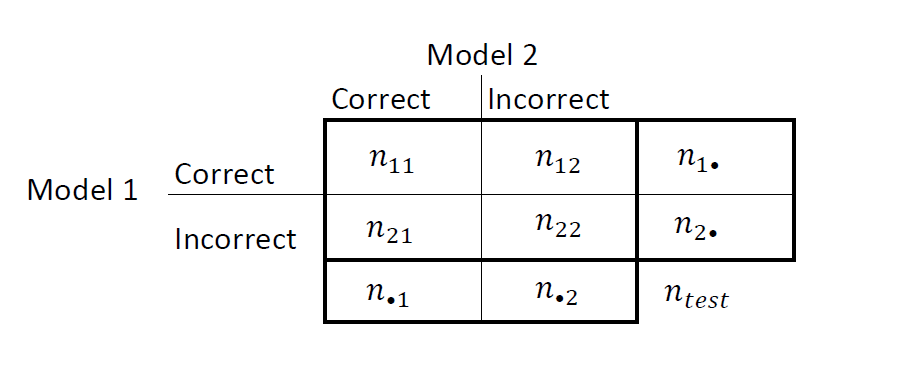
nii are the number of concordant pairs, that is, the number of observations that both models classify the same way (correctly or incorrectly). nij, i ≠ j, are the number of discordant pairs, that is, the number of observations that models classify differently (correctly or incorrectly).
The misclassification rates for Models 1 and 2 are and , respectively. A two-sided test for comparing the accuracy of the two models is
The null hypothesis suggests that the population exhibits marginal homogeneity, which reduces the null hypothesis to Also, under the null hypothesis, N12 ~ Binomial(n12 + n21,0.5) [1].
These facts are the basis for the available McNemar test variants: the asymptotic, exact-conditional, and mid-p-value McNemar tests. The definitions that follow summarize the available variants.
Asymptotic — The asymptotic McNemar test statistics and rejection regions (for significance level α) are:
For one-sided tests, the test statistic is
If where Φ is the standard Gaussian cdf, then reject H0.
For two-sided tests, the test statistic is
If , where is the χm2 cdf evaluated at x, then reject H0.
The asymptotic test requires large-sample theory, specifically, the Gaussian approximation to the binomial distribution.
The total number of discordant pairs, , must be greater than 10 ([1], Ch. 10.1.4).
In general, asymptotic tests do not guarantee nominal coverage. The observed probability of falsely rejecting the null hypothesis can exceed α, as suggested in simulation studies in [2]. However, the asymptotic McNemar test performs well in terms of statistical power.
Exact-Conditional — The exact-conditional McNemar test statistics and rejection regions (for significance level α) are ([4], [5]):
For one-sided tests, the test statistic is
If , where is the binomial cdf with sample size n and success probability p evaluated at x, then reject H0.
For two-sided tests, the test statistic is
If , then reject H0.
The exact-conditional test always attains nominal coverage. Simulation studies in [2] suggest that the test is conservative, and then show that the test lacks statistical power compared to other variants. For small or highly discrete test samples, consider using the mid-p-value test ([1], Ch. 3.6.3).
Mid-p-value test — The mid-p-value McNemar test statistics and rejection regions (for significance level α) are ([3]):
For one-sided tests, the test statistic is
If , where and are the binomial cdf and pdf, respectively, with sample size n and success probability p evaluated at x, then reject H0.
For two-sided tests, the test statistic is
If , then reject H0.
The mid-p-value test addresses the over-conservative behavior of the exact-conditional test. The simulation studies in [2] demonstrate that this test attains nominal coverage, and has good statistical power.
Classification losses indicate the accuracy of a classification model or set of predicted labels. Two classification losses are the misclassification rate and cost.
testcholdout returns the classification losses (see
e1 and e2) under the alternative hypothesis
(that is, the unrestricted classification losses).
nijk is the number of test-sample observations
with: true class k, label i assigned by the first
classification model, and label j assigned by the second classification
model. The corresponding estimated proportion is The test-set sample size is The indices are taken from 1 through K, the number of classes.
The misclassification rate, or classification error, is a scalar in the interval [0,1] representing the proportion of misclassified observations. That is, the misclassification rate for the first classification model is
For the misclassification rate of the second classification model (e2), switch the indices i and j in the formula.
Classification accuracy decreases as the misclassification rate increases to 1.
The misclassification cost is a nonnegative scalar that is a measure of classification quality relative to the values of the specified cost matrix. Its interpretation depends on the specified costs of misclassification. The misclassification cost is the weighted average of the costs of misclassification (specified in a cost matrix, C) in which the weights are the respective estimated proportions of misclassified observations. The misclassification cost for the first classification model is
where ckj is the cost of classifying an observation into class j if its true class is k. For the misclassification cost of the second classification model (e2), switch the indices i and j in the formula.
In general, for a fixed cost matrix, classification accuracy decreases as the misclassification cost increases.
Tips
It is a good practice to obtain predicted class labels by passing any trained classification model and new predictor data to the
predictmethod. For example, for predicted labels from an SVM model, seepredict.Cost-sensitive tests perform numerical optimization, which requires additional computational resources. The likelihood ratio test conducts numerical optimization indirectly by finding the root of a Lagrange multiplier in an interval. For some data sets, if the root lies close to the boundaries of the interval, then the method can fail. Therefore, if you have an Optimization Toolbox license, consider conducting the cost-sensitive chi-square test instead. For more details, see
CostTestand Cost-Sensitive Testing.
References
[1] Agresti, A. Categorical Data Analysis, 2nd Ed. John Wiley & Sons, Inc.: Hoboken, NJ, 2002.
[2] Fagerlan, M.W., S. Lydersen, and P. Laake. “The McNemar Test for Binary Matched-Pairs Data: Mid-p and Asymptotic Are Better Than Exact Conditional.” BMC Medical Research Methodology. Vol. 13, 2013, pp. 1–8.
[3] Lancaster, H.O. “Significance Tests in Discrete Distributions.” JASA, Vol. 56, Number 294, 1961, pp. 223–234.
[4] McNemar, Q. “Note on the Sampling Error of the Difference Between Correlated Proportions or Percentages.” Psychometrika, Vol. 12, Number 2, 1947, pp. 153–157.
[5] Mosteller, F. “Some Statistical Problems in Measuring the Subjective Response to Drugs.” Biometrics, Vol. 8, Number 3, 1952, pp. 220–226.
Version History
Introduced in R2015a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)