wmspca
Multiscale principal component analysis
Syntax
Description
[
returns a simplified version xsim,qual,npc_out,decsim,pca_params] = wmspca(x,level,wname,npc_in)xsim of the input matrix
x obtained from the wavelet-based multiscale principal component analysis
(PCA). The wavelet decomposition is performed using the decomposition level
level and the wavelet wname.
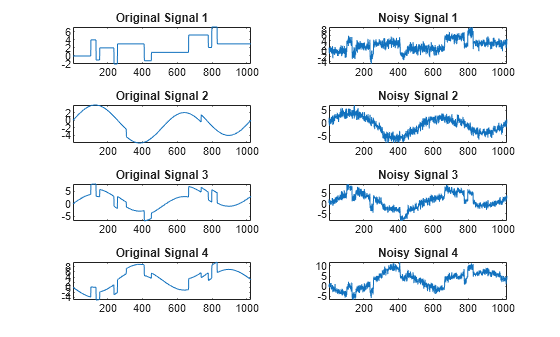
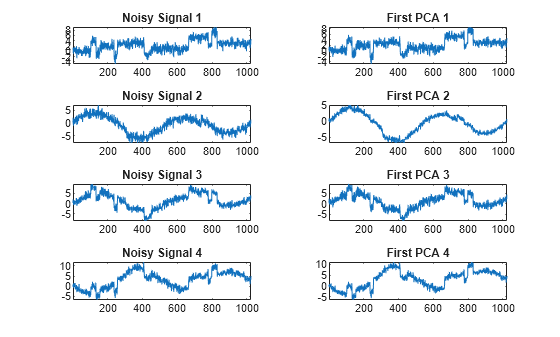
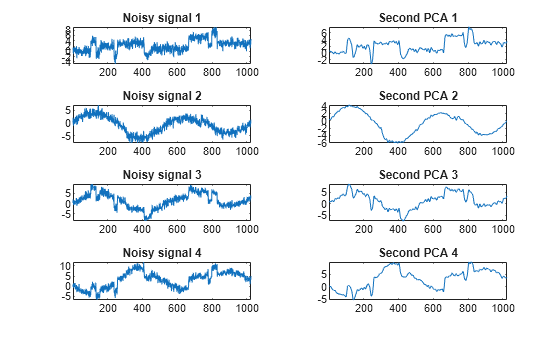
Examples
Input Arguments
Output Arguments
Algorithms
The multiscale principal components generalizes the usual PCA of a multivariate signal seen as a matrix by performing simultaneously a PCA on the matrices of details of different levels. In addition, a PCA is performed also on the coarser approximation coefficients matrix in the wavelet domain as well as on the final reconstructed matrix. By selecting conveniently the numbers of retained principal components, interesting simplified signals can be reconstructed.
References
[1] Bakshi, Bhavik R. “Multiscale PCA with Application to Multivariate Statistical Process Monitoring.” AIChE Journal 44, no. 7 (July 1998): 1596–1610. https://doi.org/10.1002/aic.690440712.
Version History
Introduced in R2006b