You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
FilterM, FiltFiltM: Fast digital filter
These functions are compatible to MATLAB's FILTER and FILTFILT commands,
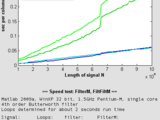
but they are faster (see screenshot):
FilterM: 30%-40% of FILTER runtime
FiltFiltM: 4%-20% of FILTFILT runtime
ADDITIONAL FEATURES:
- The dimension to operate on can be specified for FiltFiltM.
- FilterM can process the signal in backward direction. (This is the
main part of the acceleration of FiltFiltM, because it avoids to
reverse the signal two times.)
- For signals of type SINGLE, the intermediate values are stored in
DOUBLE precision to increase the accuracy. The output is converted
to SINGLE again.
- The Signal Processing Toolbox is *not* needed.
CALLING:
Y = FiltFiltM(b, a, X, Dim)
[Y, Zf] = FilterM(b, a, X, Zi, Dim, Reverse)
b, a: Filter parameters as DOUBLE vectors.
X: Signal as DOUBLE or SINGLE vector or array.
Zi, Zf: Initial and final conditions as DOUBLE or SINGLE array.
Optional, default: Zeros.
Dim: Dimension to operate on. Optional, default: 1st non-singelton.
Reverse: Flag to process the signal in reverse direction.
Optional, default: FALSE.
Y: Filtered signal, same size and type as X.
While FilterM filters in forward direction, FiltFiltM processes
the signal forward and reverse direction for a zero phase
distortion.
INTENTION:
To accelerate my FEX submission FiltFiltM, I've implemented a filter as
C-Mex, which works in reverse order. To my surprise this was faster than
running Matlab's FILTER forward, e.g. 3.7 times for a [10000 x 1] vector,
5th order Butterworth filter (Matlab 2009a, WinXP 32 bit, single core).
Therefore I've expanded the Mex such that the direction can be defined
as input. The algorithm is a direct form II transposed structure.
A future version will be mutli-threaded.
INSTALLATION:
Setup the compiler if not done before: mex -setup.
Auto-compilation: Call FilterM without inputs to start the compilation.
A pre-compiled Mex can be downloaded: http://www.n-simon.de/mex
Run the unit-tests uTest_FilterM and uTest_FiltFiltM to check validity and speed.
Tested: Matlab 6.5, 7.7, 7.8, WinXP, 32bit
Compiler: LCC2.4/3.8, BCC5.5, OWC1.8, MSVC2008
Assumed Compatibility: higher Matlab versions, Mac, Linux, 64bit
This is faster and more powerful than my former submission "FiltFiltM", which will be removed soon.
Cite As
Jan (2026). FilterM (https://www.mathworks.com/matlabcentral/fileexchange/32261-filterm), MATLAB Central File Exchange. Retrieved .
Acknowledgements
Inspired: filter1, reorganize a nD-array into matrix, and vise versa, Fast Guassian Blur
General Information
- Version 1.0.0.0 (19.9 KB)
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.0.0.0 |