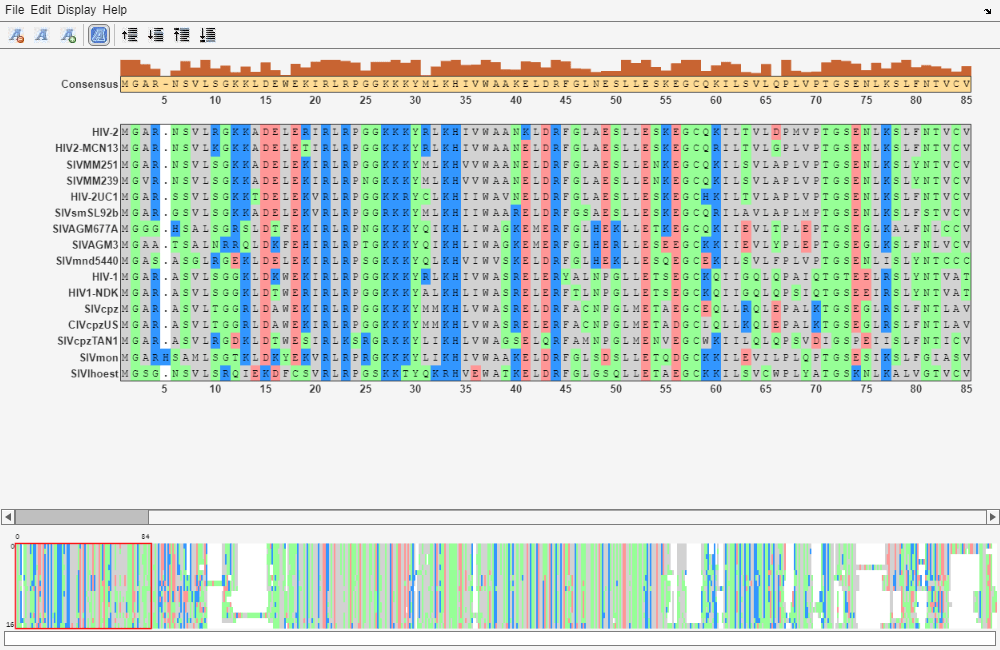
seqalignviewer
Visualize and edit multiple sequence alignment
Description
seqalignviewer opens the Sequence Alignment app, where you
can display and interactively adjust multiple sequence alignments.
seqalignviewer( loads a group
of previously multiply aligned sequences into the app, where you can view and
interactively adjust the alignment. Alignment)
seqalignviewer(
opens the app with additional options specified by one or more
Alignment,Name,Value)Name,Value pair arguments.
Tip
If gaps are available after you have selected a block from aligned sequences, then there are three regions that you can drag and move horizontally:
Selected block
Block on the left of the selection
Block on the right of the selection