Sequence Alignment
Visualize and edit multiple sequence alignments
Description
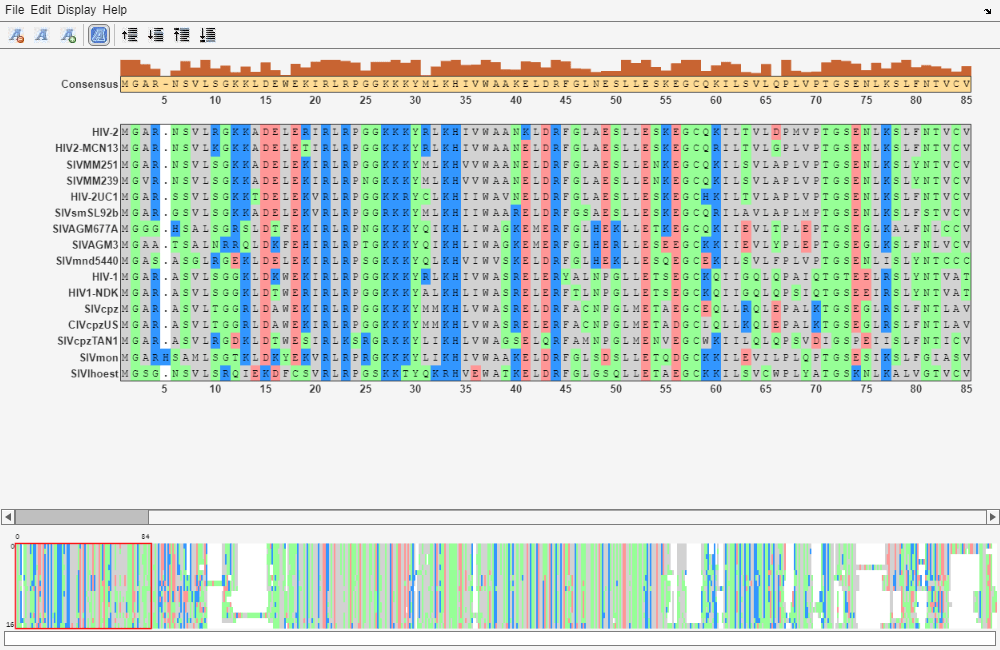
The Sequence Alignment app lets you visualize and edit multiple sequence alignments.
You can:
Inspect the sequence alignment and make manual adjustments.
View the consequence sequence information and export it to a file or MATLAB® workspace.
Generate a phylogenetic tree from aligned sequences.
Open the Sequence Alignment App
MATLAB Toolstrip: On the Apps tab, under Computational Biology, click the app icon.
MATLAB command prompt: Enter
seqalignviewer.
Programmatic Use
Version History
Introduced in R2012b