Sequence Classification Using Deep Learning
This example shows how to classify sequence data using a long short-term memory (LSTM) network.
To train a deep neural network to classify sequence data, you can use an LSTM neural network. An LSTM neural network enables you to input sequence data into a network, and make predictions based on the individual time steps of the sequence data.
This diagram illustrates sequence data flowing through a sequence classification neural network.
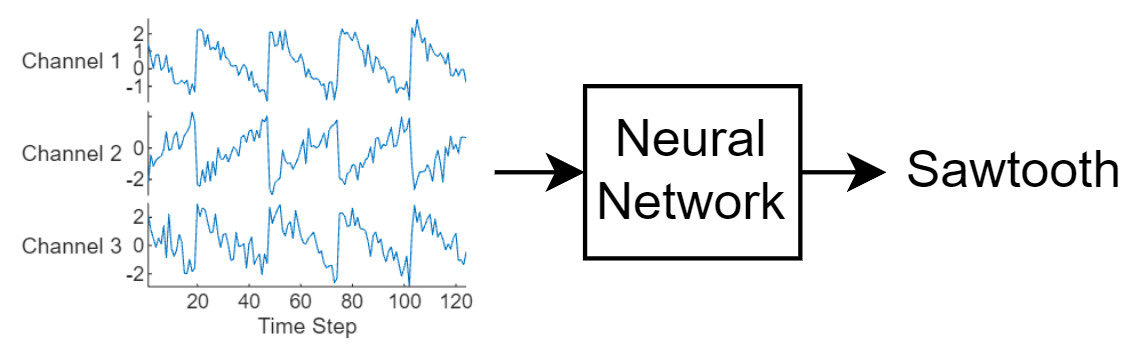
This example uses the Waveform data set. This example trains an LSTM neural network to recognize the type of waveform given time series data. The training data contains time series data for four types of waveform. Each sequence has three channels and varies in length.
Load Sequence Data
Load the example data from WaveformData. The sequence data is a numObservations-by-1 cell array of sequences, where numObservations is the number of sequences. Each sequence is a numTimeSteps-by-numChannels numeric array, where numTimeSteps is the number of time steps of the sequence and numChannels is the number of channels of the sequence. The label data is a numObservations-by-1 categorical vector.
load WaveformData Visualize some of the sequences in a plot.
numChannels = size(data{1},2);
idx = [3 4 5 12];
figure
tiledlayout(2,2)
for i = 1:4
nexttile
stackedplot(data{idx(i)},DisplayLabels="Channel "+string(1:numChannels))
xlabel("Time Step")
title("Class: " + string(labels(idx(i))))
end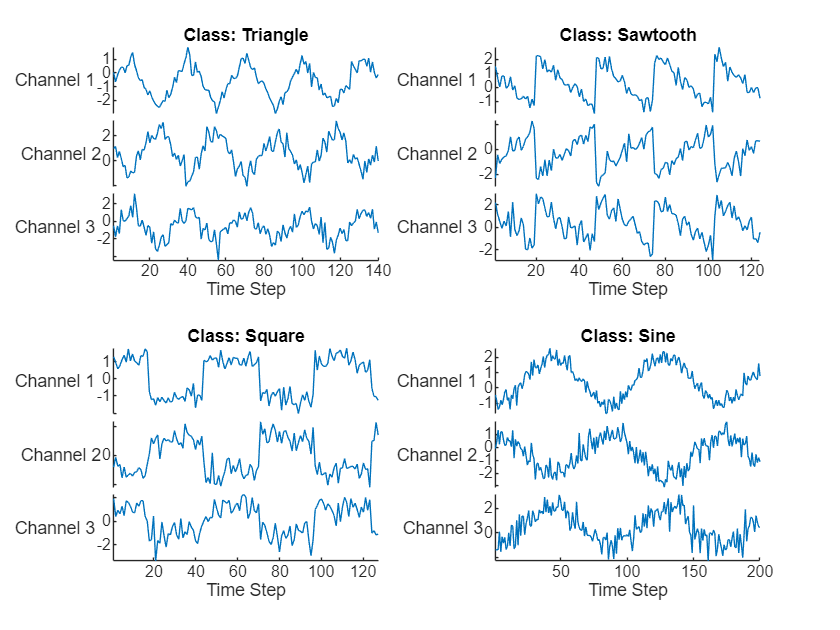
View the class names.
classNames = categories(labels)
classNames = 4×1 cell
{'Sawtooth'}
{'Sine' }
{'Square' }
{'Triangle'}
Set aside data for testing. Partition the data into a training set containing 90% of the data and a test set containing the remaining 10% of the data. To partition the data, use the trainingPartitions function, attached to this example as a supporting file. To access this file, open the example as a live script.
numObservations = numel(data); [idxTrain,idxTest] = trainingPartitions(numObservations,[0.9 0.1]); XTrain = data(idxTrain); TTrain = labels(idxTrain); XTest = data(idxTest); TTest = labels(idxTest);
Prepare Data for Padding
During training, by default, the software splits the training data into mini-batches and pads the sequences so that they have the same length. Too much padding can have a negative impact on the network performance.
To prevent the training process from adding too much padding, you can sort the training data by sequence length, and choose a mini-batch size so that sequences in a mini-batch have a similar length. The following figure shows the effect of padding sequences before and after sorting data.
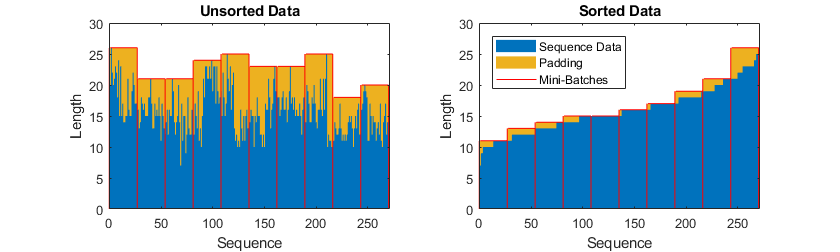
Get the sequence lengths for each observation.
numObservations = numel(XTrain); for i=1:numObservations sequence = XTrain{i}; sequenceLengths(i) = size(sequence,1); end
Sort the data by sequence length.
[sequenceLengths,idx] = sort(sequenceLengths); XTrain = XTrain(idx); TTrain = TTrain(idx);
View the sorted sequence lengths in a bar chart.
figure bar(sequenceLengths) xlabel("Sequence") ylabel("Length") title("Sorted Data")
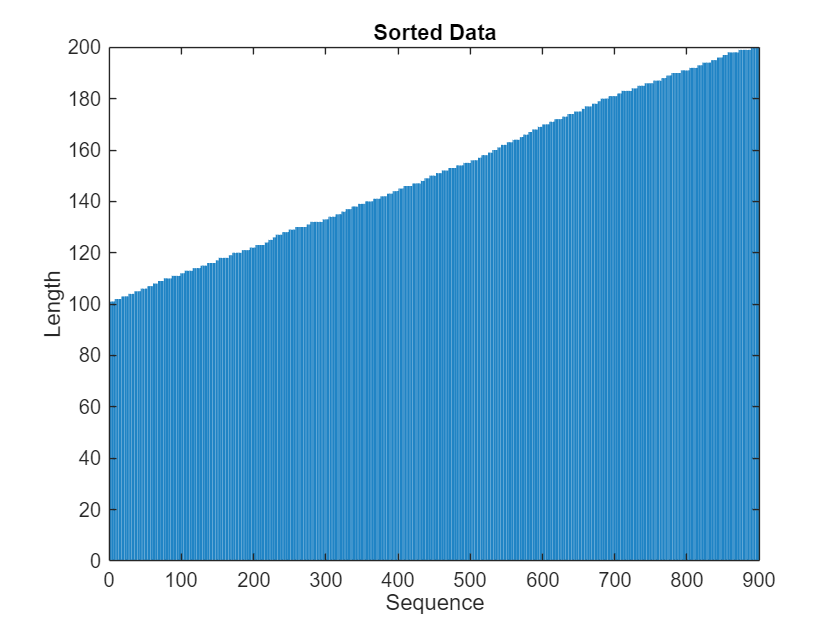
Define LSTM Neural Network Architecture
Define the LSTM neural network architecture.
Specify the input size to be the number of channels of the input data.
Specify a bidirectional LSTM layer with 120 hidden units, and output the last element of the sequence.
Finally, include a fully connected layer with an output size that matches the number of classes followed by a softmax layer.
If you have access to full sequences at prediction time, then you can use a bidirectional LSTM layer in your network. A bidirectional LSTM layer learns from the full sequence at each time step. If you do not have access to the full sequence at prediction time, for example, if you are forecasting values or predicting one time step at a time, then use an LSTM layer instead.
numHiddenUnits = 120;
numClasses = 4;
layers = [
sequenceInputLayer(numChannels)
bilstmLayer(numHiddenUnits,OutputMode="last")
fullyConnectedLayer(numClasses)
softmaxLayer]layers =
4×1 Layer array with layers:
1 '' Sequence Input Sequence input with 3 dimensions
2 '' BiLSTM BiLSTM with 120 hidden units
3 '' Fully Connected 4 fully connected layer
4 '' Softmax softmax
Specify Training Options
Specify the training options. Choosing among the options requires empirical analysis. To explore different training option configurations by running experiments, you can use the Experiment Manager app.
Train using the Adam solver.
Train for 200 epochs.
Specify a learning rate of 0.002.
Clip the gradients with a threshold of 1.
To keep the sequences sorted by length, disable shuffling.
Display the training progress in a plot and monitor the accuracy.
Disable the verbose output.
options = trainingOptions("adam", ... MaxEpochs=200, ... InitialLearnRate=0.002,... GradientThreshold=1, ... Shuffle="never", ... Plots="training-progress", ... Metrics="accuracy", ... Verbose=false);
Train LSTM Neural Network
Train the neural network using the trainnet function. For classification, use cross-entropy loss. By default, the trainnet function uses a GPU if one is available. Using a GPU requires a Parallel Computing Toolbox™ license and a supported GPU device. For information on supported devices, see GPU Computing Requirements (Parallel Computing Toolbox). Otherwise, the function uses the CPU. To specify the execution environment, use the ExecutionEnvironment training option.
net = trainnet(XTrain,TTrain,layers,"crossentropy",options);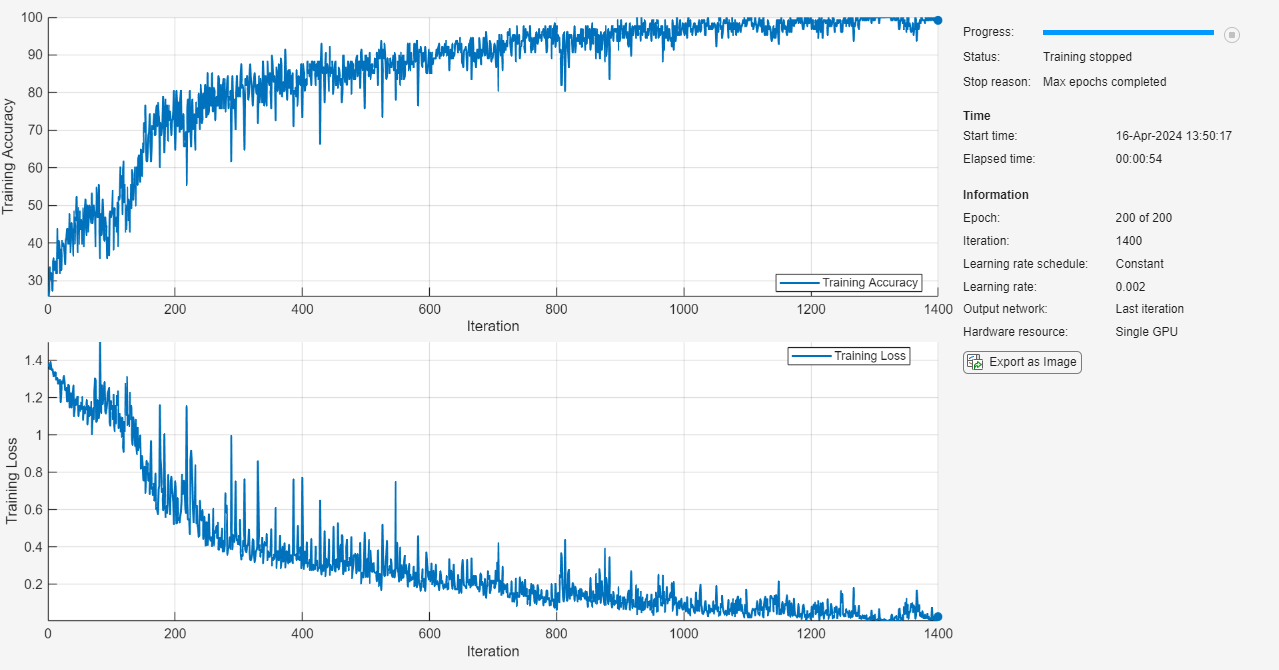
Test LSTM Neural Network
Prepare the test data. The LSTM neural network net was trained using mini-batches of sequences of similar length. Ensure that the test data is organized in the same way. Sort the test data by sequence length.
numObservationsTest = numel(XTest); for i=1:numObservationsTest sequence = XTest{i}; sequenceLengthsTest(i) = size(sequence,1); end [sequenceLengthsTest,idx] = sort(sequenceLengthsTest); XTest = XTest(idx); TTest = TTest(idx);
Test the neural network using the testnet function. For single-label classification, evaluate the accuracy. The accuracy is the percentage of correctly predicted labels. By default, the testnet function uses a GPU if one is available. To select the execution environment manually, use the ExecutionEnvironment argument of the testnet function.
acc = testnet(net,XTest,TTest,"accuracy")acc = 84
Display the classification results in a confusion chart. Make predictions using the minibatchpredict function and convert the scores to labels using the scores2label function. By default, the minibatchpredict function uses a GPU if one is available.
scores = minibatchpredict(net,XTest); YTest = scores2label(scores,classNames);
Display the classification results in a confusion chart.
figure confusionchart(TTest,YTest)
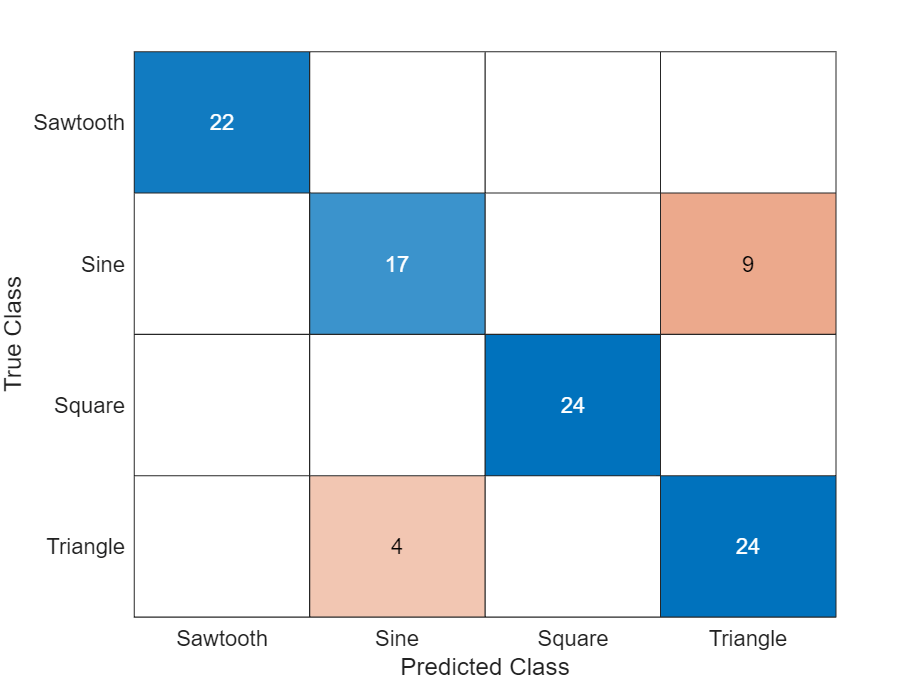
See Also
trainnet | trainingOptions | dlnetwork | lstmLayer | bilstmLayer | sequenceInputLayer
Topics
- Compare Deep Learning and ARMA Models for Time Series Modeling
- Time Series Forecasting Using Deep Learning
- Sequence-to-Sequence Classification Using Deep Learning
- Train Sequence Classification Network Using Data with Imbalanced Classes
- Sequence-to-Sequence Regression Using Deep Learning
- Sequence-to-One Regression Using Deep Learning
- Long Short-Term Memory Neural Networks
- Deep Learning in MATLAB