Train Network Using Custom Mini-Batch Datastore for Sequence Data
This example shows how to train a deep learning network on out-of-memory sequence data using a custom mini-batch datastore.
A mini-batch datastore is an implementation of a datastore with support for reading data in batches. Use mini-batch datastores to read out-of-memory data or to perform specific preprocessing operations when reading batches of data. You can use a mini-batch datastore as a source of training, validation, test, and prediction data sets for deep learning applications.
This example uses the custom mini-batch datastore sequenceDatastore attached to this example as a supporting file. You can adapt this datastore to your data by customizing the datastore functions. For an example showing how to create your own custom mini-batch datastore, see Develop Custom Mini-Batch Datastore.
Load Training Data
Load the Japanese Vowels data set as described in [1] and [2]. The zip file japaneseVowels.zip contains sequences of varying length. The sequences are divided into two folders, Train and Test, which contain training sequences and test sequences, respectively. In each of these folders, the sequences are divided into subfolders, which are numbered from 1 to 9. The names of these subfolders are the label names. A MAT file represents each sequence. Each sequence is a matrix with 12 rows, with one row for each feature, and a varying number of columns, with one column for each time step. The number of rows is the sequence dimension and the number of columns is the sequence length.
Unzip the sequence data.
filename = "japaneseVowels.zip"; outputFolder = fullfile(tempdir,"japaneseVowels"); unzip(filename,outputFolder);
Create Custom Mini-Batch Datastore
Create a custom mini-batch datastore. The mini-batch datastore sequenceDatastore reads data from a folder and gets the labels from the subfolder names.
Create a datastore containing the sequence data using sequenceDatastore.
folderTrain = fullfile(outputFolder,"Train");
dsTrain = sequenceDatastore(folderTrain)dsTrain =
sequenceDatastore with properties:
Datastore: [1×1 matlab.io.datastore.FileDatastore]
Labels: [270×1 categorical]
NumClasses: 9
SequenceDimension: 12
MiniBatchSize: 128
NumObservations: 270
Define LSTM Network Architecture
Define the LSTM network architecture. Specify the sequence dimension of the input data as the input size. Specify an LSTM layer with 100 hidden units and to output the last element of the sequence. Finally, specify a fully connected layer with output size equal to the number of classes, followed by a softmax layer.
inputSize = dsTrain.SequenceDimension;
numClasses = dsTrain.NumClasses;
numHiddenUnits = 100;
layers = [
sequenceInputLayer(inputSize)
lstmLayer(numHiddenUnits,OutputMode="last")
fullyConnectedLayer(numClasses)
softmaxLayer];Specify the training options. Specify adam as the solver and GradientThreshold as 1. Set the mini-batch size to 27 and set the maximum number of epochs to 75. To ensure that the datastore creates mini-batches of the size that the trainnet function expects, also set the mini-batch size of the datastore to the same value.
Because the mini-batches are small with short sequences, the CPU is better suited for training. Set ExecutionEnvironment to cpu. To train on a GPU, if available, set ExecutionEnvironment to "auto" (the default value).
miniBatchSize = 27; options = trainingOptions("adam", ... InputDataFormats="CTB", ... Metrics="accuracy", ... ExecutionEnvironment="cpu", ... MaxEpochs=40, ... MiniBatchSize=miniBatchSize, ... GradientThreshold=1, ... Verbose=false, ... Plots="training-progress"); dsTrain.MiniBatchSize = miniBatchSize;
Train the neural network using the trainnet function. For classification, use cross-entropy loss.
net = trainnet(dsTrain,layers,"crossentropy",options);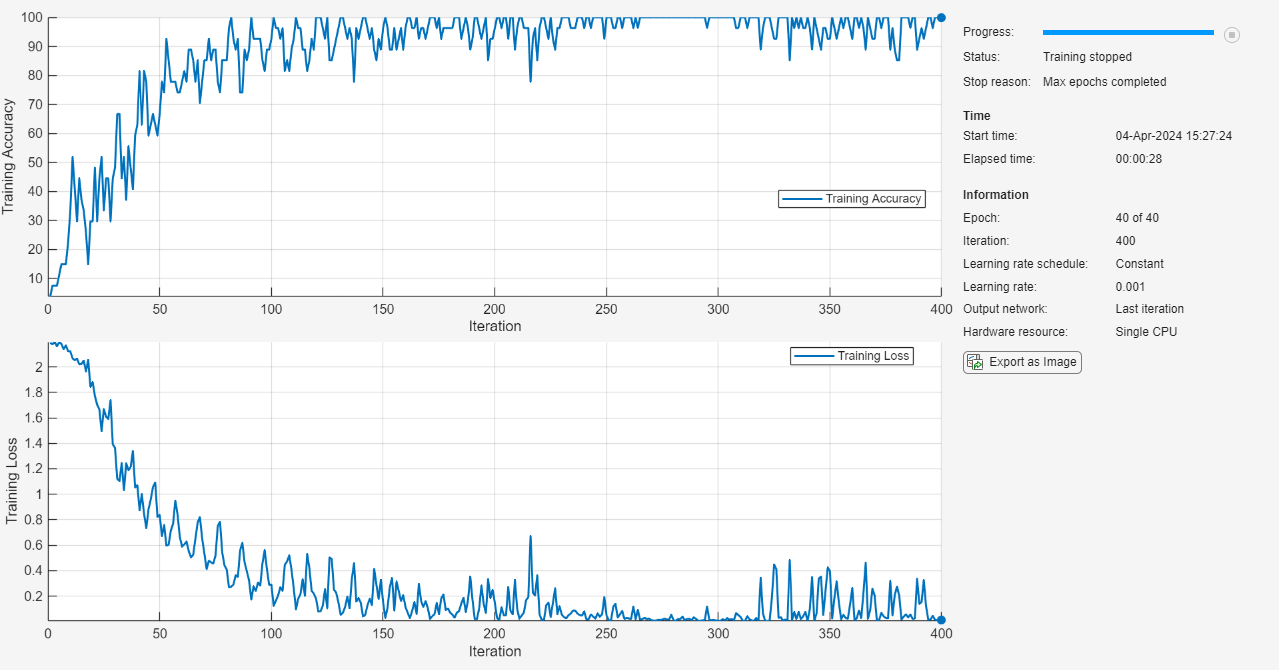
Test the Network
Create a sequence datastore from the test data. Specify the same mini-batch size as for the training data.
folderTest = fullfile(outputFolder,"Test");
dsTest = sequenceDatastore(folderTest);
dsTest.MiniBatchSize = miniBatchSize;Test the neural network using the testnet function. For single-label classification, evaluate the accuracy. The accuracy is the percentage of correct predictions. By default, the testnet function uses a GPU if one is available. To select the execution environment manually, use the ExecutionEnvironment argument of the testnet function. To ensure that the function uses mini-batches of the same size and format as for training, set the ExecutionEnvironment, InputDataFormats, and MiniBatchSize arguments to the same value as used for training.
accuracy = testnet(net,dsTest,"accuracy", ... ExecutionEnvironment="cpu", ... InputDataFormats="CTB", ... MiniBatchSize=miniBatchSize)
accuracy = 92.4324
References
[1] Kudo, M., J. Toyama, and M. Shimbo. "Multidimensional Curve Classification Using Passing-Through Regions." Pattern Recognition Letters. Vol. 20, No. 11–13, pp. 1103–1111.
[2] Kudo, M., J. Toyama, and M. Shimbo. Japanese Vowels Data Set. https://archive.ics.uci.edu/ml/datasets/Japanese+Vowels
See Also
trainnet | trainingOptions | dlnetwork | lstmLayer | sequenceInputLayer