fscnca
Feature selection using neighborhood component analysis for classification
Syntax
Description
fscnca performs feature selection using neighborhood
component analysis (NCA) for classification.
To perform NCA-based feature selection for regression, see fsrnca.
mdl = fscnca(Tbl,ResponseVarName)Tbl.
ResponseVarName is the name of the variable in
Tbl that contains the class labels.
fscnca learns the feature weights by using a diagonal
adaptation of NCA with regularization.
mdl = fscnca(X,Y,Name,Value)
Examples
Generate toy data where the response variable depends on the 3rd, 9th, and 15th predictors.
rng(0,'twister'); % For reproducibility N = 100; X = rand(N,20); y = -ones(N,1); y(X(:,3).*X(:,9)./X(:,15) < 0.4) = 1;
Fit the neighborhood component analysis model for classification.
mdl = fscnca(X,y,'Solver','sgd','Verbose',1);
o Tuning initial learning rate: NumTuningIterations = 20, TuningSubsetSize = 100
|===============================================|
| TUNING | TUNING SUBSET | LEARNING |
| ITER | FUN VALUE | RATE |
|===============================================|
| 1 | -3.755936e-01 | 2.000000e-01 |
| 2 | -3.950971e-01 | 4.000000e-01 |
| 3 | -4.311848e-01 | 8.000000e-01 |
| 4 | -4.903195e-01 | 1.600000e+00 |
| 5 | -5.630190e-01 | 3.200000e+00 |
| 6 | -6.166993e-01 | 6.400000e+00 |
| 7 | -6.255669e-01 | 1.280000e+01 |
| 8 | -6.255669e-01 | 1.280000e+01 |
| 9 | -6.255669e-01 | 1.280000e+01 |
| 10 | -6.255669e-01 | 1.280000e+01 |
| 11 | -6.255669e-01 | 1.280000e+01 |
| 12 | -6.255669e-01 | 1.280000e+01 |
| 13 | -6.255669e-01 | 1.280000e+01 |
| 14 | -6.279210e-01 | 2.560000e+01 |
| 15 | -6.279210e-01 | 2.560000e+01 |
| 16 | -6.279210e-01 | 2.560000e+01 |
| 17 | -6.279210e-01 | 2.560000e+01 |
| 18 | -6.279210e-01 | 2.560000e+01 |
| 19 | -6.279210e-01 | 2.560000e+01 |
| 20 | -6.279210e-01 | 2.560000e+01 |
o Solver = SGD, MiniBatchSize = 10, PassLimit = 5
|==========================================================================================|
| PASS | ITER | AVG MINIBATCH | AVG MINIBATCH | NORM STEP | LEARNING |
| | | FUN VALUE | NORM GRAD | | RATE |
|==========================================================================================|
| 0 | 9 | -5.658450e-01 | 4.492407e-02 | 9.290605e-01 | 2.560000e+01 |
| 1 | 19 | -6.131382e-01 | 4.923625e-02 | 7.421541e-01 | 1.280000e+01 |
| 2 | 29 | -6.225056e-01 | 3.738784e-02 | 3.277588e-01 | 8.533333e+00 |
| 3 | 39 | -6.233366e-01 | 4.947901e-02 | 5.431133e-01 | 6.400000e+00 |
| 4 | 49 | -6.238576e-01 | 3.445763e-02 | 2.946188e-01 | 5.120000e+00 |
Two norm of the final step = 2.946e-01
Relative two norm of the final step = 6.588e-02, TolX = 1.000e-06
EXIT: Iteration or pass limit reached.
Plot the selected features. The weights of the irrelevant features should be close to zero.
figure() plot(mdl.FeatureWeights,'ro') grid on xlabel('Feature index') ylabel('Feature weight')
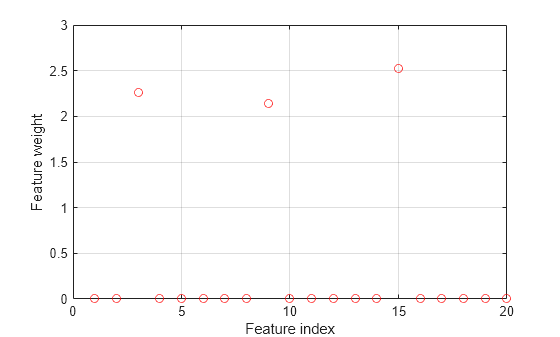
fscnca correctly detects the relevant features.
Load and partition the ovarian cancer data set, and determine if feature selection is necessary. Fit the model, plot the feature weights, and then classify observations using the selected features.
load ovariancancer;
whosName Size Bytes Class Attributes grp 216x1 26784 cell obs 216x4000 3456000 single
The obs variable consists of 216 observations with 4000 features. Each element in grp defines the group to which the corresponding row of obs belongs.
Use cvpartition to divide the data into a training set of size 160 and a test set of size 56. Both the training set and the test set have roughly the same group proportions as in grp.
rng(1,"twister"); % For reproducibility cvp = cvpartition(grp,Holdout=56)
cvp =
Hold-out cross validation partition
NumObservations: 216
NumTestSets: 1
TrainSize: 160
TestSize: 56
IsCustom: 0
IsGrouped: 0
IsStratified: 1
Properties, Methods
Xtrain = obs(cvp.training,:); ytrain = grp(cvp.training,:); Xtest = obs(cvp.test,:); ytest = grp(cvp.test,:);
To determine if feature selection is necessary, first compute the generalization error without fitting.
nca = fscnca(Xtrain,ytrain,FitMethod="none");
loss(nca,Xtest,ytest)ans = 0.0893
The software computes the generalization error of the neighborhood component analysis (NCA) feature selection model using the initial feature weights (in this case, the default feature weights) provided by fscnca.
Fit the NCA model without the regularization parameter (that is, Lambda = 0).
nca = fscnca(Xtrain,ytrain,FitMethod="exact",Lambda=0,... Solver="sgd",Standardize=true); loss(nca,Xtest,ytest)
ans = 0.0714
The improvement in the loss value suggests that feature selection is worthwhile. Tuning the regularization parameter (Lambda value) usually improves the results.
Tuning the regularization parameter for the NCA model means finding the Lambda value that produces the minimum classification loss. To tune the parameter using five-fold cross-validation:
1. Partition the training data into five folds and extract the number of validation (test) sets. For each fold, cvpartition assigns four-fifths of the data as a training set, and one-fifth of the data as a test set.
cvp = cvpartition(ytrain,KFold=5); numvalidsets = cvp.NumTestSets;
Assign Lambda values and create an array to store the loss function values.
n = length(ytrain); lambdavals = linspace(0,20,20)/n; lossvals = zeros(length(lambdavals),numvalidsets);
2. Train the NCA model for each Lambda value, using the training set in each fold.
3. Compute the classification loss for the corresponding test set in the fold using the NCA model. Record the loss value.
4. Repeat this process for all folds and all Lambda values.
for i = 1:length(lambdavals) for k = 1:numvalidsets X = Xtrain(cvp.training(k),:); y = ytrain(cvp.training(k),:); Xvalid = Xtrain(cvp.test(k),:); yvalid = ytrain(cvp.test(k),:); nca = fscnca(X,y,FitMethod="exact", ... Solver="sgd",Lambda=lambdavals(i), ... IterationLimit=30,GradientTolerance=1e-4, ... Standardize=true); lossvals(i,k) = loss(nca,Xvalid,yvalid,LossFunction="classiferror"); end end
Compute the average loss obtained from the folds for each Lambda value.
meanloss = mean(lossvals,2);
Plot the average loss values versus the Lambda values.
figure() plot(lambdavals,meanloss,"ro-") xlabel("Lambda") ylabel("Loss (MSE)") grid on
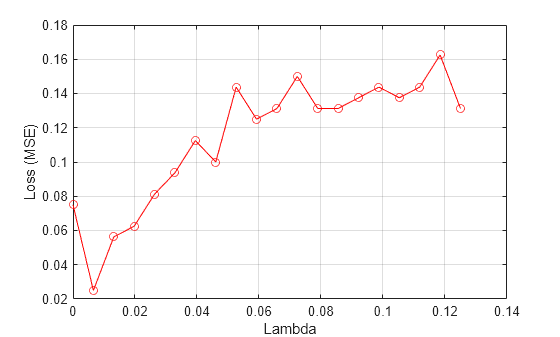
Find the best Lambda value that corresponds to the minimum average loss.
[~,idx] = min(meanloss) % Find the indexidx = 2
bestlambda = lambdavals(idx) % Find the best Lambda valuebestlambda = 0.0066
bestloss = meanloss(idx)
bestloss = 0.0312
Fit the NCA model on all the data using the best Lambda value. Use the solver sgd and standardize the predictor values.
nca = fscnca(Xtrain,ytrain,FitMethod="exact",Solver="sgd",... Lambda=bestlambda,Standardize=true,Verbose=1);
o Tuning initial learning rate: NumTuningIterations = 20, TuningSubsetSize = 100
|===============================================|
| TUNING | TUNING SUBSET | LEARNING |
| ITER | FUN VALUE | RATE |
|===============================================|
| 1 | 2.403497e+01 | 2.000000e-01 |
| 2 | 2.275050e+01 | 4.000000e-01 |
| 3 | 2.036845e+01 | 8.000000e-01 |
| 4 | 1.627647e+01 | 1.600000e+00 |
| 5 | 1.023512e+01 | 3.200000e+00 |
| 6 | 3.864283e+00 | 6.400000e+00 |
| 7 | 4.743816e-01 | 1.280000e+01 |
| 8 | -7.260138e-01 | 2.560000e+01 |
| 9 | -7.260138e-01 | 2.560000e+01 |
| 10 | -7.260138e-01 | 2.560000e+01 |
| 11 | -7.260138e-01 | 2.560000e+01 |
| 12 | -7.260138e-01 | 2.560000e+01 |
| 13 | -7.260138e-01 | 2.560000e+01 |
| 14 | -7.260138e-01 | 2.560000e+01 |
| 15 | -7.260138e-01 | 2.560000e+01 |
| 16 | -7.260138e-01 | 2.560000e+01 |
| 17 | -7.260138e-01 | 2.560000e+01 |
| 18 | -7.260138e-01 | 2.560000e+01 |
| 19 | -7.260138e-01 | 2.560000e+01 |
| 20 | -7.260138e-01 | 2.560000e+01 |
o Solver = SGD, MiniBatchSize = 10, PassLimit = 5
|==========================================================================================|
| PASS | ITER | AVG MINIBATCH | AVG MINIBATCH | NORM STEP | LEARNING |
| | | FUN VALUE | NORM GRAD | | RATE |
|==========================================================================================|
| 0 | 9 | 4.016078e+00 | 2.835465e-02 | 5.395984e+00 | 2.560000e+01 |
| 1 | 19 | -6.726156e-01 | 6.111354e-02 | 5.021138e-01 | 1.280000e+01 |
| 1 | 29 | -8.316555e-01 | 4.024186e-02 | 1.196031e+00 | 1.280000e+01 |
| 2 | 39 | -8.838656e-01 | 2.333416e-02 | 1.225834e-01 | 8.533333e+00 |
| 3 | 49 | -8.669034e-01 | 3.413162e-02 | 3.421902e-01 | 6.400000e+00 |
| 3 | 59 | -8.906936e-01 | 1.946295e-02 | 2.232511e-01 | 6.400000e+00 |
| 4 | 69 | -8.778630e-01 | 3.561290e-02 | 3.290645e-01 | 5.120000e+00 |
| 4 | 79 | -8.857135e-01 | 2.516638e-02 | 3.902979e-01 | 5.120000e+00 |
Two norm of the final step = 3.903e-01
Relative two norm of the final step = 6.171e-03, TolX = 1.000e-06
EXIT: Iteration or pass limit reached.
Plot the feature weights.
figure() plot(nca.FeatureWeights,"ro") xlabel("Feature Index") ylabel("Feature Weight") grid on
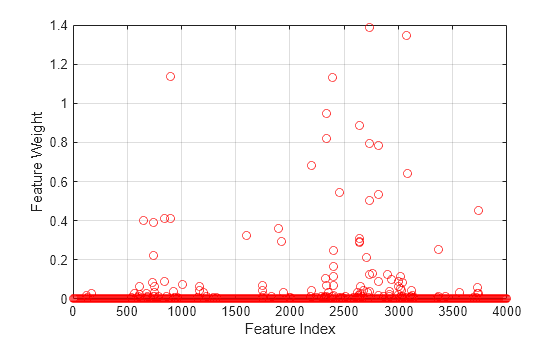
Most of the feature weights are very close to zero, which means that they are irrelevant. Some features have much higher feature weight values. In this case, to select a reasonable number of predictors, specify a threshold of 0.02 times the maximum feature weight value.
selidx = find(nca.FeatureWeights > 0.02*max(1,max(nca.FeatureWeights)))
selidx = 72×1
565
611
654
681
737
743
744
750
754
839
840
897
899
925
1010
⋮
Compute the classification loss using the test set.
loss(nca,Xtest,ytest)
ans = 0.0179
Extract the features with feature weights greater than the specified threshold value from the training data.
features = Xtrain(:,selidx);
Apply a support vector machine classifier to the reduced training set using the selected features.
svmMdl = fitcsvm(features,ytrain);
Evaluate the accuracy of the trained classifier on the test data, which has not been used for feature selection.
loss(svmMdl,Xtest(:,selidx),ytest)
ans = single
0
Input Arguments
Sample data used to train the model, specified as a table. Each row of Tbl corresponds to one observation, and each column corresponds to one predictor variable.
Data Types: table
Response variable name, specified as the name of a variable in Tbl. The
remaining variables in the table are predictors.
Data Types: char | string
Predictor variable values, specified as an n-by-p matrix, where n is the number of observations and p is the number of predictor variables.
Data Types: single | double
Explanatory model of the response variable and a subset of the predictor variables, specified
as a string or a character vector in the form "Y~x1+x2+x3". In this
form, Y represents the response variable, and x1,
x2, and x3 represent the predictor
variables.
To specify a subset of variables in Tbl as predictors for training the model, use a formula. If you specify a formula, then the software does not use any variables in Tbl that do not appear in formula.
The variable names in the formula must be both variable names in Tbl
(Tbl.Properties.VariableNames) and valid MATLAB® identifiers. You can verify the variable names in Tbl
by using the isvarname function. If the variable
names are not valid, then you can convert them by using the matlab.lang.makeValidName function.
Data Types: char | string
Class labels, specified as a categorical array, logical vector, numeric vector, string
array, cell array of character vectors of length n, or character
matrix with n rows. n is the number of
observations. Element i or row i of
Y is the class label corresponding to row i
of X (observation i).
Data Types: single | double | logical | char | string | cell | categorical
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: 'Solver','sgd','Weights',W,'Lambda',0.0003 specifies
the solver as the stochastic gradient descent, the observation weights
as the values in the vector W, and sets the regularization
parameter at 0.0003.
Fitting Options
Method for fitting the model, specified as the comma-separated
pair consisting of 'FitMethod' and one of the following:
'exact'— Performs fitting using all of the data.'none'— No fitting. Use this option to evaluate the generalization error of the NCA model using the initial feature weights supplied in the call to fscnca.'average'— Divides the data into partitions (subsets), fits each partition using theexactmethod, and returns the average of the feature weights. You can specify the number of partitions using theNumPartitionsname-value pair argument.
Example: 'FitMethod','none'
Number of partitions to split the data for using with 'FitMethod','average' option,
specified as the comma-separated pair consisting of 'NumPartitions' and
an integer value between 2 and n, where n is
the number of observations.
Example: 'NumPartitions',15
Data Types: double | single
Regularization parameter to prevent overfitting, specified as the
comma-separated pair consisting of 'Lambda' and a
nonnegative scalar.
As the number of observations n increases, the chance of overfitting decreases and the required amount of regularization also decreases. See Identify Relevant Features for Classification and Tune Regularization Parameter to Detect Features Using NCA for Classification to learn how to tune the regularization parameter.
Example: 'Lambda',0.002
Data Types: double | single
Width of the kernel, specified as the comma-separated pair consisting
of 'LengthScale' and a positive real scalar.
A length scale value of 1 is sensible when all predictors are
on the same scale. If the predictors in X are
of very different magnitudes, then consider standardizing the predictor
values using 'Standardize',true and setting 'LengthScale',1.
Example: 'LengthScale',1.5
Data Types: double | single
Categorical predictors list, specified as one of the values in this table.
| Value | Description |
|---|---|
| Vector of positive integers | Each entry in the vector is an index value corresponding to the column of the predictor data (X) that contains a categorical variable. |
| Logical vector | A true entry means that the corresponding column of predictor data (X) is a categorical variable. |
| Character matrix | Each row of the matrix is the name of a predictor variable in the table X. The names must match the entries in PredictorNames. Pad the names with extra blanks so each row of the character matrix has the same length. |
| String array or cell array of character vectors | Each element in the array is the name of a predictor variable in the table X. The names must match the entries in PredictorNames. |
"all" | All predictors are categorical. |
By default, if the predictor data is in a table,
fscnca assumes that a variable is categorical if it is a
logical vector, categorical vector, character array, string array, or cell array of
character vectors. If the predictor data is a matrix, fscnca
assumes that all predictors are continuous. To identify any other predictors as
categorical predictors, specify them by using the
CategoricalPredictors name-value argument.
For the identified categorical predictors, fscnca creates dummy variables using two different schemes, depending on whether a categorical variable is unordered or ordered:
For an unordered categorical variable,
fscncacreates one dummy variable for each level of the categorical variable.For an ordered categorical variable,
fscncacreates one less dummy variable than the number of categories. For details, see Automatic Creation of Dummy Variables.
For the table X, categorical predictors can be ordered and unordered. For the matrix X, fscnca treats categorical predictors as unordered.
Example: CategoricalPredictors="all"
Data Types: double | logical | char | string
Predictor variable names, specified as a string array of unique names or cell array of unique
character vectors. The functionality of PredictorNames depends on the
way you supply the training data.
If you supply
Xas a matrix, then you can usePredictorNamesto assign names to the predictor variables inX.The order of the names in
PredictorNamesmust correspond to the predictor order inX. That is,PredictorNames{1}is the name ofX(:,1),PredictorNames{2}is the name ofX(:,2), and so on. Also,size(X,2)andnumel(PredictorNames)must be equal.By default,
PredictorNamesis{'X1','X2',...}.
If you supply
Xas a table, then you can usePredictorNamesto specify which predictor variables to use in training. That is,fscncauses only the predictor variables inPredictorNamesand the response variable during training.PredictorNamesmust be a subset ofX.Properties.VariableNamesand cannot include the name of the response variable.By default,
PredictorNamescontains the names of all predictor variables.Specify the predictors for training using either
PredictorNamesor a formula string inY(such as'y ~ x1 + x2 + x3'), but not both.
Example: "PredictorNames={"SepalLength","SepalWidth","PetalLength","PetalWidth"}
Data Types: string | cell
Response variable name, specified as a character vector or string scalar.
If you supply
Y, then you can useResponseNameto specify a name for the response variable.If you supply
ResponseVarNameorformula, then you cannot useResponseName.
Example: ResponseName="response"
Data Types: char | string
Initial feature weights, specified as an M-by-1 vector of positive numbers,
where M is the number of predictor variables after dummy variables
are created for categorical variables (for details, see
CategoricalPredictors).
The regularized objective function for optimizing feature weights is nonconvex. As a result,
using different initial feature weights might give different results. Setting all
initial feature weights to 1 generally works well, but in some cases, random
initialization using rand(M,1) might give better quality
solutions.
For more information about feature weights, see Neighborhood Component Analysis (NCA) Feature Selection.
Data Types: double | single
Observation weights, specified as the comma-separated pair consisting of
'Weights' and an n-by-1 vector of real
positive scalars. Use observation weights to specify higher importance of some
observations compared to others. The default weights assign equal importance to all
observations.
Data Types: double | single
Prior probabilities for each class, specified as the comma-separated pair consisting of
'Prior' and one of the following:
'empirical'—fscncaobtains the prior class probabilities from class frequencies.'uniform'—fscncasets all class probabilities equal.Structure with two fields:
ClassProbs— Vector of class probabilities. If these are numeric values with a total greater than 1,fsncanormalizes them to add up to 1.ClassNames— Class names corresponding to the class probabilities inClassProbs.
Example: 'Prior','uniform'
Indicator for standardizing the predictor data, specified as the comma-separated pair
consisting of 'Standardize' and either false or
true. For more information, see Impact of Standardization.
Example: 'Standardize',true
Data Types: logical
Verbosity level indicator for the convergence summary display,
specified as the comma-separated pair consisting of 'Verbose' and
one of the following:
0 — No convergence summary
1 — Convergence summary, including norm of gradient and objective function values
> 1 — More convergence information, depending on the fitting algorithm
When using
'minibatch-lbfgs'solver and verbosity level > 1, the convergence information includes iteration the log from intermediate mini-batch LBFGS fits.
Example: 'Verbose',1
Data Types: double | single
Solver type for estimating feature weights, specified as the
comma-separated pair consisting of 'Solver' and
one of the following:
'lbfgs'— Limited memory Broyden-Fletcher-Goldfarb-Shanno (LBFGS) algorithm'sgd'— Stochastic gradient descent (SGD) algorithm'minibatch-lbfgs'— Stochastic gradient descent with LBFGS algorithm applied to mini-batches
Default is 'lbfgs' for n ≤
1000, and 'sgd' for n > 1000.
Example: 'solver','minibatch-lbfgs'
Loss function, specified as the comma-separated pair consisting
of 'LossFunction' and one of the following.
'classiferror'— Misclassification error@— Custom loss function handle. A loss function has this form.lossfunfunction L = lossfun(Yu,Yv) % calculation of loss ...
Yuis a u-by-1 vector andYvis a v-by-1 vector.Lis a u-by-v matrix of loss values such thatL(i,j)is the loss value forYu(i)andYv(j).
The objective function for minimization includes the loss function l(yi,yj) as follows:
where w is the feature weight vector, n is the number of observations, and p is the number of predictor variables. pij is the probability that xj is the reference point for xi. For details, see NCA Feature Selection for Classification.
Example: 'LossFunction',@lossfun
Memory size, in MB, to use for objective function and gradient
computation, specified as the comma-separated pair consisting of 'CacheSize' and
an integer.
Example: 'CacheSize',1500MB
Data Types: double | single
LBFGS Options
Size of history buffer for Hessian approximation for the 'lbfgs' solver,
specified as the comma-separated pair consisting of 'HessianHistorySize' and
a positive integer. At each iteration the function uses the most recent HessianHistorySize iterations
to build an approximation to the inverse Hessian.
Example: 'HessianHistorySize',20
Data Types: double | single
Initial step size for the 'lbfgs' solver,
specified as the comma-separated pair consisting of 'InitialStepSize' and
a positive real scalar. By default, the function determines the initial
step size automatically.
Data Types: double | single
Line search method, specified as the comma-separated pair consisting
of 'LineSearchMethod' and one of the following:
'weakwolfe'— Weak Wolfe line search'strongwolfe'— Strong Wolfe line search'backtracking'— Backtracking line search
Example: 'LineSearchMethod','backtracking'
Maximum number of line search iterations, specified as the comma-separated
pair consisting of 'MaxLineSearchIterations' and
a positive integer.
Example: 'MaxLineSearchIterations',25
Data Types: double | single
Relative convergence tolerance on the gradient norm for solver lbfgs,
specified as the comma-separated pair consisting of 'GradientTolerance' and
a positive real scalar.
Example: 'GradientTolerance',0.000002
Data Types: double | single
SGD Options
Initial learning rate for the 'sgd' solver,
specified as the comma-separated pair consisting of 'InitialLearningRate' and
a positive real scalar.
When using solver type 'sgd', the learning
rate decays over iterations starting with the value specified for 'InitialLearningRate'.
The default 'auto' means that the initial
learning rate is determined using experiments on small subsets of
data. Use the NumTuningIterations name-value
pair argument to specify the number of iterations for automatically
tuning the initial learning rate. Use the TuningSubsetSize name-value
pair argument to specify the number of observations to use for automatically
tuning the initial learning rate.
For solver type 'minibatch-lbfgs', you can
set 'InitialLearningRate' to a very high value.
In this case, the function applies LBFGS to each mini-batch separately
with initial feature weights from the previous mini-batch.
To make sure the chosen initial learning rate decreases the
objective value with each iteration, plot the Iteration versus
the Objective values saved in the mdl.FitInfo property.
You can use the refit method with 'InitialFeatureWeights' equal
to mdl.FeatureWeights to start from the current
solution and run additional iterations
Example: 'InitialLearningRate',0.9
Data Types: double | single
Number of observations to use in each batch for the 'sgd' solver,
specified as the comma-separated pair consisting of 'MiniBatchSize' and
a positive integer from 1 to n.
Example: 'MiniBatchSize',25
Data Types: double | single
Maximum number of passes through all n observations
for solver 'sgd', specified as the comma-separated
pair consisting of 'PassLimit' and a positive integer.
Each pass through all of the data is called an epoch.
Example: 'PassLimit',10
Data Types: double | single
Frequency of batches for displaying convergence summary for
the 'sgd' solver , specified as the comma-separated
pair consisting of 'NumPrint' and a positive integer.
This argument applies when the 'Verbose' value
is greater than 0. NumPrint mini-batches are
processed for each line of the convergence summary that is displayed
on the command line.
Example: 'NumPrint',5
Data Types: double | single
Number of tuning iterations for the 'sgd' solver,
specified as the comma-separated pair consisting of 'NumTuningIterations' and
a positive integer. This option is valid only for 'InitialLearningRate','auto'.
Example: 'NumTuningIterations',15
Data Types: double | single
Number of observations to use for tuning the initial learning
rate, specified as the comma-separated pair consisting of 'TuningSubsetSize' and
a positive integer value from 1 to n. This option
is valid only for 'InitialLearningRate','auto'.
Example: 'TuningSubsetSize',25
Data Types: double | single
SGD or LBFGS Options
Maximum number of iterations, specified as the comma-separated
pair consisting of 'IterationLimit' and a positive
integer. The default is 10000 for SGD and 1000 for LBFGS and mini-batch
LBFGS.
Each pass through a batch is an iteration. Each pass through all of the data is an epoch. If the data is divided into k mini-batches, then every epoch is equivalent to k iterations.
Example: 'IterationLimit',250
Data Types: double | single
Convergence tolerance on the step size, specified as the comma-separated
pair consisting of 'StepTolerance' and a positive
real scalar. The 'lbfgs' solver uses an absolute
step tolerance, and the 'sgd' solver uses a relative
step tolerance.
Example: 'StepTolerance',0.000005
Data Types: double | single
Mini-Batch LBFGS Options
Maximum number of iterations per mini-batch LBFGS step, specified
as the comma-separated pair consisting of 'MiniBatchLBFGSIterations' and
a positive integer.
Example: 'MiniBatchLBFGSIterations',15
Data Types: double | single
Note
The mini-batch LBFGS algorithm is a combination of SGD and LBFGS methods. Therefore, all of the name-value pair arguments that apply to SGD and LBFGS solvers also apply to the mini-batch LBFGS algorithm.
Output Arguments
Neighborhood component analysis model for classification, returned
as a FeatureSelectionNCAClassification object.
Version History
Introduced in R2016b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)