plotDiagnostics
Plot observation diagnostics of generalized linear regression model
Syntax
Description
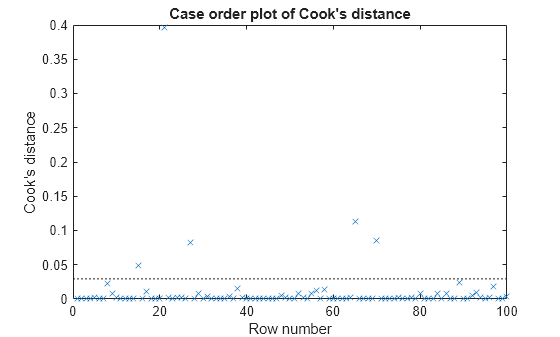
plotDiagnostics creates a plot of observation
diagnostics, such as leverage and Cook's distance, to identify outliers and influential
observations.
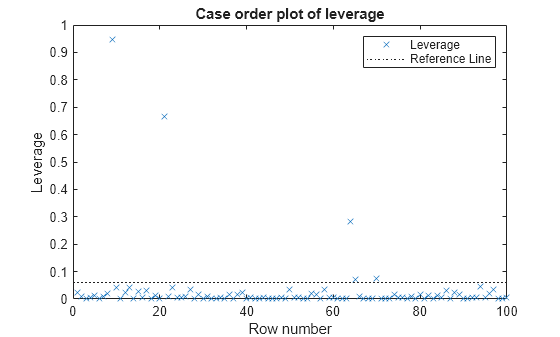
plotDiagnostics( creates a leverage
plot of the generalized linear regression model (mdl)mdl)
observations. A dotted line in the plot represents the recommended threshold
values.
plotDiagnostics(___,
specifies additional options using one or more name-value pair arguments in addition
to any of the input argument combinations in the previous syntaxes. For example, you
can specify the marker symbol and size for the data points.Name,Value)
h = plotDiagnostics(___)h to modify the properties of a specific line or contour
after you create the plot. For a list of properties, see Line Properties and Contour Properties.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
More About
Tips
The data cursor displays the values of the selected plot point in a data tip (small text box located next to the data point). The data tip includes the x-axis and y-axis values for the selected point, along with the observation name or number.
Use
legend('show')to show the pre-populated legend.
Alternative Functionality
A GeneralizedLinearModel object provides multiple plotting functions.
When verifying a model, use
plotDiagnosticsto find questionable data and to understand the effect of each observation. Also, useplotResidualsto analyze the residuals of the model.After fitting a model, use
plotPartialDependenceto understand the effect of a particular predictor. Also, useplotSliceto plot slices through the prediction surface.
References
[1] Neter, J., M. H. Kutner, C. J. Nachtsheim, and W. Wasserman. Applied Linear Statistical Models, Fourth Edition. Chicago: McGraw-Hill Irwin, 1996.