residuals
Residuals of fitted linear mixed-effects model
Description
R = residuals(lme,Name,Value)lme with
additional options specified by one or more Name,Value pair
arguments.
For example, you can specify Pearson or standardized residuals, or residuals with contributions from only fixed effects.
Examples
Load the sample data.
load('weight.mat');weight contains data from a longitudinal study, where 20 subjects are randomly assigned to 4 exercise programs, and their weight loss is recorded over six 2-week time periods. This is simulated data.
Store the data in a table. Define Subject and Program as categorical variables.
tbl = table(InitialWeight,Program,Subject,Week,y); tbl.Subject = nominal(tbl.Subject); tbl.Program = nominal(tbl.Program);
Fit a linear mixed-effects model where the initial weight, type of program, week, and the interaction between the week and type of program are the fixed effects. The intercept and week vary by subject.
lme = fitlme(tbl,'y ~ InitialWeight + Program*Week + (Week|Subject)');Compute the fitted values and raw residuals.
F = fitted(lme); R = residuals(lme);
Plot the residuals versus the fitted values.
plot(F,R,'bx') xlabel('Fitted Values') ylabel('Residuals')
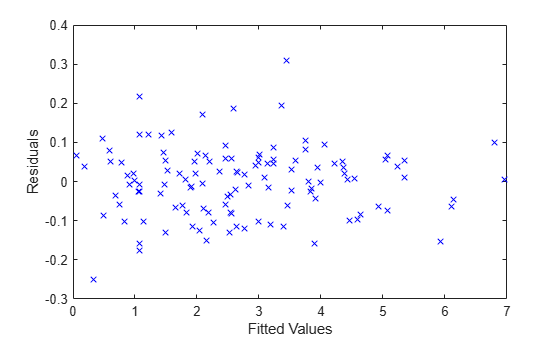
Now, plot the residuals versus the fitted values, grouped by program.
figure(); gscatter(F,R,Program)
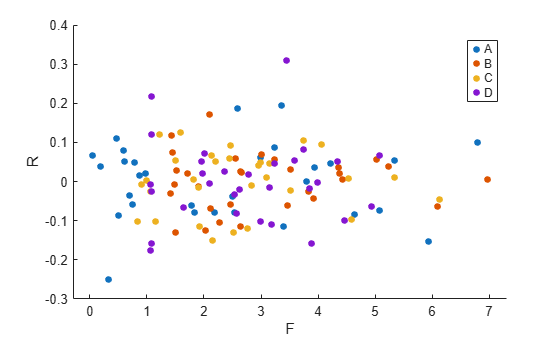
The residuals seem to behave similarly across levels of the program as expected.
Load the sample data.
load carbigStore the variables for miles per gallon (MPG), acceleration, horsepower, cylinders, and model year in a table.
tbl = table(MPG,Acceleration,Horsepower,Cylinders,Model_Year);
Fit a linear mixed-effects model for miles per gallon (MPG), with fixed effects for acceleration, horsepower, and the cylinders, and potentially correlated random effects for intercept and acceleration grouped by model year.
lme = fitlme(tbl,'MPG ~ Acceleration + Horsepower + Cylinders + (Acceleration|Model_Year)');Compute the conditional Pearson residuals and display the first five residuals.
PR = residuals(lme,'ResidualType','Pearson'); PR(1:5)
ans = 5×1
-0.0533
0.0652
0.3655
-0.0106
-0.3340
Compute the marginal Pearson residuals and display the first five residuals.
PRM = residuals(lme,'ResidualType','Pearson','Conditional',false); PRM(1:5)
ans = 5×1
-0.1250
0.0130
0.3242
-0.0861
-0.3006
Load the sample data.
load carbigStore the variables for miles per gallon (MPG), acceleration, horsepower, cylinders, and model year in a table.
tbl = table(MPG,Acceleration,Horsepower,Cylinders,Model_Year);
Fit a linear mixed-effects model for miles per gallon (MPG), with fixed effects for acceleration, horsepower, and the cylinders, and potentially correlated random effects for intercept and acceleration grouped by model year.
lme = fitlme(tbl,'MPG ~ Acceleration + Horsepower + Cylinders + (Acceleration|Model_Year)');Draw a histogram of the raw residuals with a normal fit.
r = residuals(lme); histfit(r)
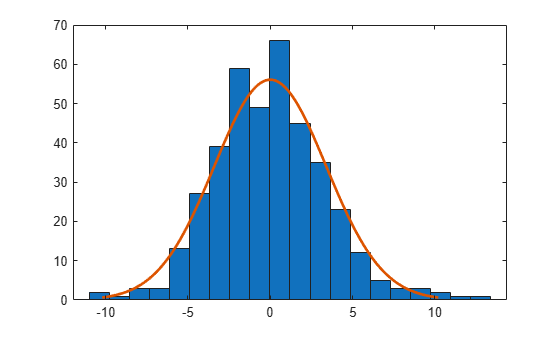
Normal distribution seems to be a good fit for the residuals.
Compute the conditional Pearson and standardized residuals and create box plots of all three types of residuals.
pr = residuals(lme,'ResidualType','Pearson'); st = residuals(lme,'ResidualType','Standardized'); X = [r pr st]; boxplot(X)
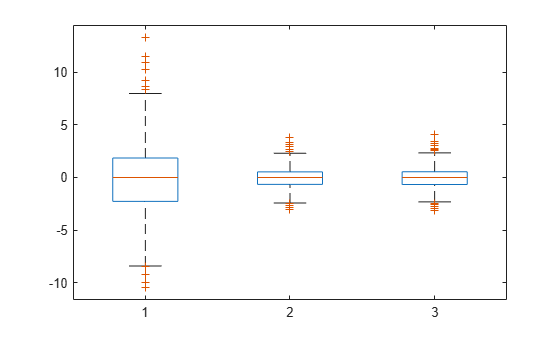
Red plus signs show the observations with residuals above or below and , where and are the 25th and 75th percentiles, respectively.
Find the observations with residuals that are 2.5 standard deviations above and below the mean.
find(r > mean(r,'omitnan') + 2.5*std(r,'omitnan'))
ans = 7×1
62
252
255
330
337
341
396
find(r < mean(r,'omitnan') - 2.5*std(r,'omitnan'))
ans = 3×1
119
324
375
Input Arguments
Linear mixed-effects model, specified as a LinearMixedModel object constructed using fitlme or fitlmematrix.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: R =
residuals(lme,'ResidualType','Pearson');
Indicator for conditional residuals, specified as the comma-separated
pair consisting of 'Conditional' and one of the
following.
True | Contribution from both fixed effects and random effects (conditional) |
False | Contribution from only fixed effects (marginal) |
Example: 'Conditional,'False'
Residual type, specified as one of the following.
| Residual Type | Conditional | Marginal |
|---|---|---|
"Raw" |
|
|
"Pearson" |
|
|
"Standardized" |
|
|
Example: ResidualType="Standardized"
Data Types: string | char
Output Arguments
Residuals of the fitted linear mixed-effects model lme returned as an
n-by-1 vector, where n is the
number of observations.
More About
Conditional residuals include contributions from both fixed and random effects, whereas marginal residuals include contribution from only fixed effects.
Suppose the linear mixed-effects model lmehas
an n-by-p fixed-effects design
matrix X and an n-by-q random-effects
design matrix Z. Also, suppose the p-by-1
estimated fixed-effects vector is , and the q-by-1
estimated best linear unbiased predictor (BLUP) vector of random effects
is . The fitted conditional response
is
and the fitted marginal response is
residuals can return three types of residuals:
raw, Pearson, and standardized. For any type, you can compute the
conditional or the marginal residuals. For example, the conditional
raw residual is
and the marginal raw residual is
For more information on other types of residuals, see the ResidualType name-value
pair argument.
Version History
Introduced in R2013b
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)