What Is Biomedical Signal Processing?
Biomedical signal processing involves acquiring and preprocessing physiological signals and extracting meaningful information to identify patterns and trends within the signals.
Sources of biomedical signals include neural activity, cardiac rhythm, muscle movement, and other physiological activities. Signals such as electrocardiogram (ECG), electroencephalogram (EEG), electromyography (EMG) can be captured non-invasively and used for diagnosis and as indicators of overall health.
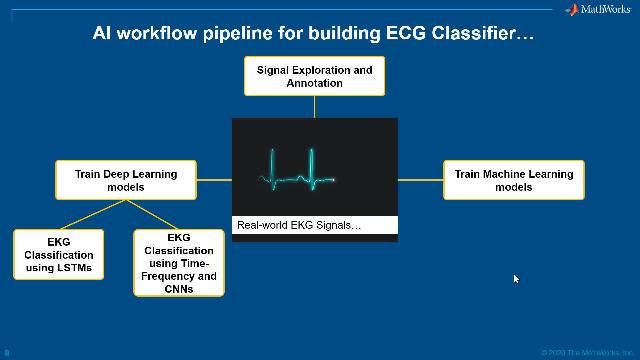
The biomedical signal processing workflow involves:
- Signal Acquisition
- Signal Visualization and Annotation
- Artifact Removal and Preprocessing
- Feature Extraction
The extracted features are then fed into classification models or used directly for diagnosis.

Workflow for processing biomedical signals.
MATLAB® provides many signal processing capabilities for this workflow, especially for signal preprocessing and feature extraction.
Signal Acquisition: With MATLAB, you can interface with hardware equipment to acquire physiological signals. For instance, with the Raspberry Pi and Arduino Support Packages, you can interface with embedded boards like Raspberry Pi, Arduino, and EKGShield to collect data from these sensors. You can also access and analyze signals stored in files such as from EDF, Excel® files and MAT-files.
Signal Visualization and Annotation: MATLAB provides built-in apps to help you analyze and visualize signals in time, frequency, and time-frequency domains without writing any code. These capabilities can help you in understanding which preprocessing techniques you can use to enhance the information contained within signals.
You can also annotate the biomedical signals and prepare them for downstream workflows such as machine learning and deep learning using the Signal Labeler app. Signals can be labeled manually or using algorithms like those that find peaks and transition points.

Analyzing an ECG signal in time, frequency and time-frequency domains with the Signal Analyzer app in MATLAB.
Artifact Removal and Signal Filtering: Biomedical signals often contains noise or unwanted artifacts that can distort the analysis of the signals. For instance, while measuring EKG signals, activities such as breathing and walking can add unwanted components. One of the main challenges in preprocessing biomedical signals is to remove unwanted artifacts while preserving the sharp features within signals. Most popular techniques for artifact removal are digital filtering, adaptive filtering, independent component analysis (ICA), and recursive least square. A combination of preprocessing techniques may also be used to address the limitations of individual techniques.

Digital filter used for signal preprocessing.
Feature Extraction with Signal Processing: Feature extraction can be accomplished manually or automatically. Signal processing techniques like AR modeling, Fourier analysis, and spectral estimation can be used to manually compute key features from signals. Time-frequency transformations, such as the short-time Fourier transform (STFT) can be used as signal representations for training data in machine learning and deep learning models. Automatic feature extraction techniques like wavelet scattering can be used to reduce dimensionality and extract important features. These features can be used directly for diagnosis or as input to machine learning and deep learning classifiers.

Time-frequency analysis used to extract features from ECG signals for classification.
After developing the signal processing algorithms, you can prototype designs and create proofs-of-concept by automatically generating real-time code. You can also generate VHDL and Verilog code to deploy signal processing algorithms on devices such as FPGAs and ASICs in order to prototype and build low-power medical devices. MATLAB Coder™ and Embedded Coder™ allow you to generate optimized C/C++ code to deploy the algorithms on embedded hardware. High-performance GPUs can be used to speed up preprocessing, feature extraction, and model inference.
For more details, see Signal Processing Toolbox™ and Wavelet Toolbox™
Examples and How To
Customer Stories
Examples
Software Reference
See also: DSP System Toolbox™, Statistics and Machine Learning Toolbox™, embedded code generation, GPUs for signal processing algorithms in MATLAB