incrementalLearner
Convert binary classification support vector machine (SVM) model to incremental learner
Description
IncrementalMdl = incrementalLearner(Mdl)IncrementalMdl, using the traditionally trained linear SVM model
object or SVM model template object in Mdl.
If you specify a traditionally trained model, then its property values reflect the
knowledge gained from Mdl (parameters and hyperparameters of the
model). Therefore, IncrementalMdl can predict labels given new
observations, and it is warm, meaning that its predictive performance
is tracked.
IncrementalMdl = incrementalLearner(Mdl,Name,Value)IncrementalMdl before its
predictive performance is tracked. For example,
'MetricsWarmupPeriod',50,'MetricsWindowSize',100 specifies a preliminary
incremental training period of 50 observations before performance metrics are tracked, and
specifies processing 100 observations before updating the window performance metrics.
Examples
Train an SVM model by using fitcsvm, and then convert it to an incremental learner.
Load and Preprocess Data
Load the human activity data set.
load humanactivityFor details on the data set, enter Description at the command line.
Responses can be one of five classes: Sitting, Standing, Walking, Running, or Dancing. Dichotomize the response by identifying whether the subject is moving (actid > 2).
Y = actid > 2;
Train SVM Model
Fit an SVM model to the entire data set. Discard the support vectors (Alpha) from the model so that the software uses the linear coefficients (Beta) for prediction.
TTMdl = fitcsvm(feat,Y); TTMdl = discardSupportVectors(TTMdl)
TTMdl =
ClassificationSVM
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: [0 1]
ScoreTransform: 'none'
NumObservations: 24075
Beta: [60×1 double]
Bias: -6.4280
KernelParameters: [1×1 struct]
BoxConstraints: [24075×1 double]
ConvergenceInfo: [1×1 struct]
IsSupportVector: [24075×1 logical]
Solver: 'SMO'
Properties, Methods
TTMdl is a ClassificationSVM model object representing a traditionally trained SVM model.
Convert Trained Model
Convert the traditionally trained SVM model to a binary classification linear model for incremental learning.
IncrementalMdl = incrementalLearner(TTMdl)
IncrementalMdl =
incrementalClassificationLinear
IsWarm: 1
Metrics: [1×2 table]
ClassNames: [0 1]
ScoreTransform: 'none'
Beta: [60×1 double]
Bias: -6.4280
Learner: 'svm'
Properties, Methods
IncrementalMdl is an incrementalClassificationLinear model object prepared for incremental learning using SVM.
The
incrementalLearnerfunction Initializes the incremental learner by passing learned coefficients to it, along with other informationTTMdlextracted from the training data.IncrementalMdlis warm (IsWarmis1), which means that incremental learning functions can start tracking performance metrics.The
incrementalLearnerfunction specifies to train the model using the adaptive scale-invariant solver, whereasfitcsvmtrainedTTMdlusing theSMOsolver.
Predict Responses
An incremental learner created from converting a traditionally trained model can generate predictions without further processing.
Predict classification scores for all observations using both models.
[~,ttscores] = predict(TTMdl,feat); [~,ilcores] = predict(IncrementalMdl,feat); compareScores = norm(ttscores(:,1) - ilcores(:,1))
compareScores = 0
The difference between the scores generated by the models is 0.
The default solver is the adaptive scale-invariant solver. If you specify this solver, you do not need to tune any parameters for training. However, if you specify either the standard SGD or ASGD solver instead, you can also specify an estimation period, during which the incremental fitting functions tune the learning rate.
Load the human activity data set.
load humanactivityFor details on the data set, enter Description at the command line.
Responses can be one of five classes: Sitting, Standing, Walking, Running, and Dancing. Dichotomize the response by identifying whether the subject is moving (actid > 2).
Y = actid > 2;
Randomly split the data in half: the first half for training a model traditionally, and the second half for incremental learning.
n = numel(Y); rng(1) % For reproducibility cvp = cvpartition(n,'Holdout',0.5); idxtt = training(cvp); idxil = test(cvp); % First half of data Xtt = feat(idxtt,:); Ytt = Y(idxtt); % Second half of data Xil = feat(idxil,:); Yil = Y(idxil);
Fit an SVM model to the first half of the data. Standardize the predictor data by setting 'Standardize',true.
TTMdl = fitcsvm(Xtt,Ytt,'Standardize',true);The Mu and Sigma properties of TTMdl contain the predictor data sample means and standard deviations, respectively.
Suppose that the distribution of the predictors is not expected to change in the future. Convert the traditionally trained SVM model to a binary classification linear model for incremental learning. Specify the standard SGD solver and an estimation period of 2000 observations (the default is 1000 when a learning rate is required).
IncrementalMdl = incrementalLearner(TTMdl,'Solver','sgd','EstimationPeriod',2000);
IncrementalMdl is an incrementalClassificationLinear model object. Because the predictor data of TTMdl is standardized (TTMdl.Mu and TTMdl.Sigma are nonempty), incrementalLearner prepares incremental learning functions to standardize supplied predictor data by using the previously learned moments (stored in IncrementalMdl.Mu and IncrementalMdl.Sigma).
Fit the incremental model to the second half of the data by using the fit function. At each iteration:
Simulate a data stream by processing 10 observations at a time.
Overwrite the previous incremental model with a new one fitted to the incoming observations.
Store the initial learning rate and to see how the coefficients and rate evolve during training.
% Preallocation nil = numel(Yil); numObsPerChunk = 10; nchunk = floor(nil/numObsPerChunk); learnrate = [IncrementalMdl.LearnRate; zeros(nchunk,1)]; beta1 = [IncrementalMdl.Beta(1); zeros(nchunk,1)]; % Incremental fitting for j = 1:nchunk ibegin = min(nil,numObsPerChunk*(j-1) + 1); iend = min(nil,numObsPerChunk*j); idx = ibegin:iend; IncrementalMdl = fit(IncrementalMdl,Xil(idx,:),Yil(idx)); beta1(j + 1) = IncrementalMdl.Beta(1); learnrate(j + 1) = IncrementalMdl.LearnRate; end
IncrementalMdl is an incrementalClassificationLinear model object trained on all the data in the stream.
To see how the initial learning rate and evolve during training, plot them on separate tiles.
t = tiledlayout(2,1); nexttile plot(beta1) ylabel('\beta_1') xline(IncrementalMdl.EstimationPeriod/numObsPerChunk,'r-.') nexttile plot(learnrate) ylabel('Initial Learning Rate') xline(IncrementalMdl.EstimationPeriod/numObsPerChunk,'r-.') xlabel(t,'Iteration')
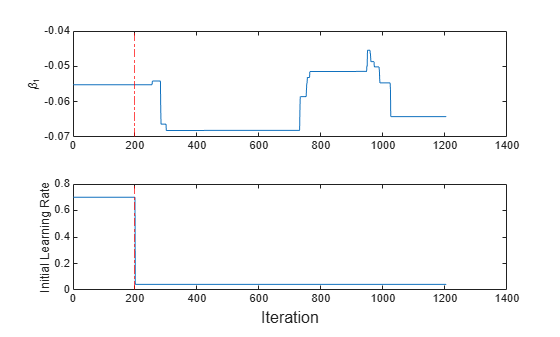
The initial learning rate jumps from 0.7 to its autotuned value after the estimation period. During training, the software uses a learning rate that gradually decays from the initial value specified in the LearnRateSchedule property of IncrementalMdl.
Because fit does not fit the model to the streaming data during the estimation period, is constant for the first 200 iterations (2000 observations). Then, changes during incremental fitting.
Use a trained SVM model to initialize an incremental learner. Prepare the incremental learner by specifying a metrics warm-up period, during which the updateMetricsAndFit function only fits the model. Specify a metrics window size of 500 observations.
Load the human activity data set.
load humanactivityFor details on the data set, enter Description at the command line
Responses can be one of five classes: Sitting, Standing, Walking, Running, and Dancing. Dichotomize the response by identifying whether the subject is moving (actid > 2).
Y = actid > 2;
Because the data set is grouped by activity, shuffle it to reduce bias. Then, randomly split the data in half: the first half for training a model traditionally, and the second half for incremental learning.
n = numel(Y); rng(1) % For reproducibility cvp = cvpartition(n,'Holdout',0.5); idxtt = training(cvp); idxil = test(cvp); shuffidx = randperm(n); X = feat(shuffidx,:); Y = Y(shuffidx); % First half of data Xtt = X(idxtt,:); Ytt = Y(idxtt); % Second half of data Xil = X(idxil,:); Yil = Y(idxil);
Fit an SVM model to the first half of the data.
TTMdl = fitcsvm(Xtt,Ytt);
Convert the traditionally trained SVM model to a binary classification linear model for incremental learning. Specify the following:
A performance metrics warm-up period of 2000 observations
A metrics window size of 500 observations
Use of classification error and hinge loss to measure the performance of the model
IncrementalMdl = incrementalLearner(TTMdl,'MetricsWarmupPeriod',2000,'MetricsWindowSize',500,... 'Metrics',["classiferror" "hinge"]);
Fit the incremental model to the second half of the data by using the updateMetricsAndFit function. At each iteration:
Simulate a data stream by processing 20 observations at a time.
Overwrite the previous incremental model with a new one fitted to the incoming observations.
Store , the cumulative metrics, and the window metrics to see how they evolve during incremental learning.
% Preallocation nil = numel(Yil); numObsPerChunk = 20; nchunk = ceil(nil/numObsPerChunk); ce = array2table(zeros(nchunk,2),'VariableNames',["Cumulative" "Window"]); hinge = array2table(zeros(nchunk,2),'VariableNames',["Cumulative" "Window"]); beta1 = [IncrementalMdl.Beta(1); zeros(nchunk,1)]; % Incremental fitting for j = 1:nchunk ibegin = min(nil,numObsPerChunk*(j-1) + 1); iend = min(nil,numObsPerChunk*j); idx = ibegin:iend; IncrementalMdl = updateMetricsAndFit(IncrementalMdl,Xil(idx,:),Yil(idx)); ce{j,:} = IncrementalMdl.Metrics{"ClassificationError",:}; hinge{j,:} = IncrementalMdl.Metrics{"HingeLoss",:}; beta1(j + 1) = IncrementalMdl.Beta(1); end
IncrementalMdl is an incrementalClassificationLinear model object trained on all the data in the stream. During incremental learning and after the model is warmed up, updateMetricsAndFit checks the performance of the model on the incoming observations, and then fits the model to those observations.
To see how the performance metrics and evolve during training, plot them on separate tiles.
t = tiledlayout(3,1); nexttile plot(beta1) ylabel('\beta_1') xlim([0 nchunk]); xline(IncrementalMdl.MetricsWarmupPeriod/numObsPerChunk,'r-.'); nexttile h = plot(ce.Variables); xlim([0 nchunk]); ylabel('Classification Error') xline(IncrementalMdl.MetricsWarmupPeriod/numObsPerChunk,'r-.'); legend(h,ce.Properties.VariableNames,'Location','northwest') nexttile h = plot(hinge.Variables); xlim([0 nchunk]); ylabel('Hinge Loss') xline(IncrementalMdl.MetricsWarmupPeriod/numObsPerChunk,'r-.'); legend(h,hinge.Properties.VariableNames,'Location','northwest') xlabel(t,'Iteration')
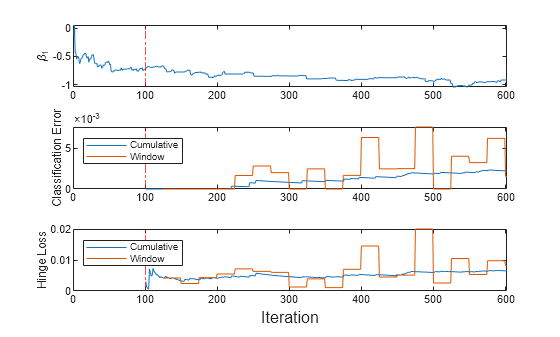
The plot suggests that updateMetricsAndFit does the following:
Fit during all incremental learning iterations.
Compute the performance metrics after the metrics warm-up period only.
Compute the cumulative metrics during each iteration.
Compute the window metrics after processing 500 observations (25 iterations).
Input Arguments
Traditionally trained linear SVM model or SVM model template, specified as a model object returned by its training or processing function.
| Model Object or Template Object | Training or Processing Function |
|---|---|
ClassificationSVM model
object | fitcsvm |
CompactClassificationSVM model
object | fitcsvm or compact |
| SVM model template object | templateSVM |
Note
Incremental learning functions support only numeric input predictor data. If
Mdlwas trained on categorical data, you must prepare an encoded version of the categorical data to use incremental learning functions. Usedummyvarto convert each categorical variable to a numeric matrix of dummy variables. Then, concatenate all dummy variable matrices and any other numeric predictors, in the same way that the training function encodes categorical data. For more details, see Dummy Variables.If
Mdlis a SVM model template object,incrementalLearnerdetermines whether to standardize the predictor variables based on theStandardizeproperty of the model template object. For more information, see Standardize Data.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: 'Solver','scale-invariant','MetricsWindowSize',100 specifies
the adaptive scale-invariant solver for objective optimization, and specifies processing 100
observations before updating the window performance metrics.
General Options
Objective function minimization technique, specified as a value in this table.
| Value | Description | Notes |
|---|---|---|
'scale-invariant' | Adaptive scale-invariant solver for incremental learning [1] |
|
'sgd' | Stochastic gradient descent (SGD) [3][2] |
|
'asgd' | Average stochastic gradient descent (ASGD) [4] |
|
The linear model for incremental learning (IncrementalMdl) does
not support the solver used to train the traditionally trained linear SVM model
Mdl or the solver specified in the SVM model template object
Mdl. By default, the incrementalLearner function
sets IncrementalMdl to use the adaptive scale-invariant solver
('scale-invariant').
Example: 'Solver','sgd'
Data Types: char | string
Number of observations processed by the incremental model to estimate hyperparameters before training or tracking performance metrics, specified as the comma-separated pair consisting of 'EstimationPeriod' and a nonnegative integer.
Note
If
Mdlis prepared for incremental learning (all hyperparameters required for training are specified),incrementalLearnerforcesEstimationPeriodto0.If
Mdlis not prepared for incremental learning,incrementalLearnersetsEstimationPeriodto1000.
For more details, see Estimation Period.
Example: 'EstimationPeriod',100
Data Types: single | double
SGD and ASGD Solver Options
Mini-batch size, specified as the comma-separated pair consisting of
'BatchSize' and a positive integer. At each learning cycle during
training, incrementalLearner uses BatchSize observations to
compute the subgradient.
The number of observations for the last mini-batch (last learning cycle in each function
call of fit or updateMetricsAndFit) can be
smaller than BatchSize. For example, if you supply 25 observations to
fit or updateMetricsAndFit, the function uses
10 observations for the first two learning cycles and uses 5 observations for the last
learning cycle.
Example: 'BatchSize',1
Data Types: single | double
Ridge (L2) regularization term strength, specified as the comma-separated pair consisting of 'Lambda' and a nonnegative scalar.
Example: 'Lambda',0.01
Data Types: single | double
Initial learning rate, specified as the comma-separated pair consisting of
'LearnRate' and 'auto' or a positive scalar.
LearnRate controls the optimization step size by scaling the
objective subgradient.
The learning rate controls the optimization step size by scaling the objective
subgradient. LearnRate specifies an initial value for the learning
rate, and LearnRateSchedule determines
the learning rate for subsequent learning cycles.
When you specify 'auto':
The initial learning rate is
0.7.If
EstimationPeriod>0,fitandupdateMetricsAndFitchange the rate to1/sqrt(1+max(sum(X.^2,obsDim)))at the end ofEstimationPeriod. When the observations are the columns of the predictor dataXcollected during the estimation period, theobsDimvalue is1; otherwise, the value is2.
Example: 'LearnRate',0.001
Data Types: single | double | char | string
Learning rate schedule, specified as the comma-separated pair consisting of 'LearnRateSchedule' and a value in this table, where LearnRate specifies the initial learning rate ɣ0.
| Value | Description |
|---|---|
'constant' | The learning rate is ɣ0 for all learning cycles. |
'decaying' | The learning rate at learning cycle t is
|
Example: 'LearnRateSchedule','constant'
Data Types: char | string
Adaptive Scale-Invariant Solver Options
Flag for shuffling the observations in the batch at each iteration, specified as the comma-separated pair consisting of 'Shuffle' and a value in this table.
| Value | Description |
|---|---|
true | The software shuffles an incoming chunk of data before the
fit function fits the model. This action
reduces bias induced by the sampling scheme. |
false | The software processes the data in the order received. |
Example: 'Shuffle',false
Data Types: logical
Performance Metrics Options
Model performance metrics to track during incremental learning with the updateMetrics or updateMetricsAndFit function, specified as a built-in loss function name, string vector of names, function handle (@metricName), structure array of function handles, or cell vector of names, function handles, or structure arrays.
The following table lists the built-in loss function names. You can specify more than one by using a string vector.
| Name | Description |
|---|---|
"binodeviance" | Binomial deviance |
"classiferror" | Classification error |
"exponential" | Exponential loss |
"hinge" | Hinge loss |
"logit" | Logistic loss |
"quadratic" | Quadratic loss |
For more details on the built-in loss functions, see loss.
Example: 'Metrics',["classiferror" "hinge"]
To specify a custom function that returns a performance metric, use function handle notation. The function must have this form:
metric = customMetric(C,S)
The output argument
metricis an n-by-1 numeric vector, where each element is the loss of the corresponding observation in the data processed by the incremental learning functions during a learning cycle.You specify the function name (
customMetric).Cis an n-by-2 logical matrix with rows indicating the class to which the corresponding observation belongs. The column order corresponds to the class order in the model for incremental learning. CreateCby settingC(=p,q)1, if observationpqp0.Sis an n-by-2 numeric matrix of predicted classification scores.Sis similar to thescoreoutput ofpredict, where rows correspond to observations in the data, and the column order corresponds to the class order in the model for incremental learning.S(is the classification score of observationp,q)pq
To specify multiple custom metrics and assign a custom name to each, use a structure array. To specify a combination of built-in and custom metrics, use a cell vector.
Example: 'Metrics',struct('Metric1',@customMetric1,'Metric2',@customMetric2)
Example: 'Metrics',{@customMetric1 @customMetric2 'logit' struct('Metric3',@customMetric3)}
updateMetrics and updateMetricsAndFit store specified metrics in a table in the property IncrementalMdl.Metrics. The data type of Metrics determines the row names of the table.
'Metrics' Value Data Type | Description of Metrics Property Row Name | Example |
|---|---|---|
| String or character vector | Name of corresponding built-in metric | Row name for "classiferror" is "ClassificationError" |
| Structure array | Field name | Row name for struct('Metric1',@customMetric1) is "Metric1" |
| Function handle to function stored in a program file | Name of function | Row name for @customMetric is "customMetric" |
| Anonymous function | CustomMetric_, where Metrics | Row name for @(C,S)customMetric(C,S)... is CustomMetric_1 |
For more details on performance metrics options, see Performance Metrics.
Data Types: char | string | struct | cell | function_handle
Number of observations the incremental model must be fit to before it tracks
performance metrics in its Metrics property, specified as a
nonnegative integer. The incremental model is warm after incremental fitting functions
fit (EstimationPeriod + MetricsWarmupPeriod)
observations to the incremental model.
For more details on performance metrics options, see Performance Metrics.
Example: 'MetricsWarmupPeriod',50
Data Types: single | double
Number of observations to use to compute window performance metrics, specified as a positive integer.
For more details on performance metrics options, see Performance Metrics.
Example: 'MetricsWindowSize',100
Data Types: single | double
Output Arguments
Binary classification linear model for incremental learning, returned as an incrementalClassificationLinear model object. IncrementalMdl is also configured to generate predictions given new data (see predict).
If you specify a traditionally trained model object in
Mdl,incrementalLearnerpasses the values of theMdlproperties to corresponding properties ofIncrementalMdlto initializeIncrementalMdlfor incremental learning.Property Description BetaScaled linear model coefficients, Mdl.Beta/Mdl.KernelParameters.Scale, a numeric vectorBiasModel intercept, a numeric scalar ClassNamesClass labels for binary classification, two-element list MuPredictor variable means, a numeric vector NumPredictorsNumber of predictors, a positive integer PriorPrior class label distribution, a numeric vector SigmaPredictor variable standard deviations, a numeric vector ScoreTransformScore transformation function, a function name or function handle Note that
incrementalLearnerdoes not use theCostproperty of the traditionally trained model inMdlbecauseincrementalClassificationLineardoes not support this property.
More About
Incremental learning, or online learning, is a branch of machine learning concerned with processing incoming data from a data stream, possibly given little to no knowledge of the distribution of the predictor variables, aspects of the prediction or objective function (including tuning parameter values), or whether the observations are labeled. Incremental learning differs from traditional machine learning, where enough labeled data is available to fit to a model, perform cross-validation to tune hyperparameters, and infer the predictor distribution.
Given incoming observations, an incremental learning model processes data in any of the following ways, but usually in this order:
Predict labels.
Measure the predictive performance.
Check for structural breaks or drift in the model.
Fit the model to the incoming observations.
For more details, see Incremental Learning Overview.
The adaptive scale-invariant solver for incremental learning, introduced in [1], is a gradient-descent-based objective solver for training linear predictive models. The solver is hyperparameter free, insensitive to differences in predictor variable scales, and does not require prior knowledge of the distribution of the predictor variables. These characteristics make it well suited to incremental learning.
The standard SGD and ASGD solvers are sensitive to differing scales among the predictor variables, resulting in models that can perform poorly. To achieve better accuracy using SGD and ASGD, you can standardize the predictor data, and tune the regularization and learning rate parameters. For traditional machine learning, enough data is available to enable hyperparameter tuning by cross-validation and predictor standardization. However, for incremental learning, enough data might not be available (for example, observations might be available only one at a time) and the distribution of the predictors might be unknown. These characteristics make parameter tuning and predictor standardization difficult or impossible to do during incremental learning.
The incremental fitting functions for classification fit and updateMetricsAndFit use the more aggressive ScInOL2 version of the algorithm.
Algorithms
During the estimation period, the incremental fitting functions fit and updateMetricsAndFit use the
first incoming EstimationPeriod observations
to estimate (tune) hyperparameters required for incremental training. Estimation occurs only
when EstimationPeriod is positive. This table describes the
hyperparameters and when they are estimated, or tuned.
| Hyperparameter | Model Property | Usage | Conditions |
|---|---|---|---|
| Predictor means and standard deviations |
| Standardize predictor data | The hyperparameters are estimated when both of these conditions apply:
|
| Learning rate | LearnRate | Adjust the solver step size | The hyperparameter is estimated when both of these conditions apply:
|
During the estimation period, fit does not fit the model, and updateMetricsAndFit does not fit the model or update the performance metrics. At the end of the estimation period, the functions update the properties that store the hyperparameters.
If incremental learning functions are configured to standardize predictor variables, they do so using the means and standard deviations stored in the Mu and Sigma properties of the incremental learning model IncrementalMdl.
If you standardize the predictor data when you train the input model
Mdlby usingfitcsvm, the following conditions apply:incrementalLearnerpasses the means inMdl.Muand standard deviations inMdl.Sigmato the corresponding incremental learning model properties.Incremental learning functions always standardize the predictor data.
When you set
Standardize=trueby using theStandardizename-value argument oftemplateSVM, and theMdl.MuandMdl.Sigmaproperties are empty, the following conditions apply:If the estimation period is positive (see the
EstimationPeriodproperty ofIncrementalMdl), incremental fitting functions estimate the means and standard deviations using the estimation period observations.If the estimation period is 0,
incrementalLearnerforces the estimation period to1000. Consequently, incremental fitting functions estimate new predictor variable means and standard deviations during the forced estimation period.
When incremental fitting functions estimate predictor means and standard deviations, the functions compute weighted means and weighted standard deviations using the estimation period observations. Specifically, the functions standardize predictor j (xj) using
xj is predictor j, and xjk is observation k of predictor j in the estimation period.
pk is the prior probability of class k (
Priorproperty of the incremental model).wj is observation weight j.
The
updateMetricsandupdateMetricsAndFitfunctions are incremental learning functions that track model performance metrics ('Metrics') from new data when the incremental model is warm (IsWarmproperty). An incremental model becomes warm afterfitorupdateMetricsAndFitfit the incremental model to'MetricsWarmupPeriod'observations, which is the metrics warm-up period.If
'EstimationPeriod'> 0, the functions estimate hyperparameters before fitting the model to data. Therefore, the functions must process an additionalEstimationPeriodobservations before the model starts the metrics warm-up period.The
Metricsproperty of the incremental model stores two forms of each performance metric as variables (columns) of a table,CumulativeandWindow, with individual metrics in rows. When the incremental model is warm,updateMetricsandupdateMetricsAndFitupdate the metrics at the following frequencies:Cumulative— The functions compute cumulative metrics since the start of model performance tracking. The functions update metrics every time you call the functions and base the calculation on the entire supplied data set.Window— The functions compute metrics based on all observations within a window determined by the'MetricsWindowSize'name-value pair argument.'MetricsWindowSize'also determines the frequency at which the software updatesWindowmetrics. For example, ifMetricsWindowSizeis 20, the functions compute metrics based on the last 20 observations in the supplied data (X((end – 20 + 1):end,:)andY((end – 20 + 1):end)).Incremental functions that track performance metrics within a window use the following process:
Store a buffer of length
MetricsWindowSizefor each specified metric, and store a buffer of observation weights.Populate elements of the metrics buffer with the model performance based on batches of incoming observations, and store corresponding observation weights in the weights buffer.
When the buffer is filled, overwrite
IncrementalMdl.Metrics.Windowwith the weighted average performance in the metrics window. If the buffer is overfilled when the function processes a batch of observations, the latest incomingMetricsWindowSizeobservations enter the buffer, and the earliest observations are removed from the buffer. For example, supposeMetricsWindowSizeis 20, the metrics buffer has 10 values from a previously processed batch, and 15 values are incoming. To compose the length 20 window, the functions use the measurements from the 15 incoming observations and the latest 5 measurements from the previous batch.
The software omits an observation with a
NaNscore when computing theCumulativeandWindowperformance metric values.
References
[1] Kempka, Michał, Wojciech Kotłowski, and Manfred K. Warmuth. "Adaptive Scale-Invariant Online Algorithms for Learning Linear Models." Preprint, submitted February 10, 2019. https://arxiv.org/abs/1902.07528.
[2] Langford, J., L. Li, and T. Zhang. “Sparse Online Learning Via Truncated Gradient.” J. Mach. Learn. Res., Vol. 10, 2009, pp. 777–801.
[3] Shalev-Shwartz, S., Y. Singer, and N. Srebro. “Pegasos: Primal Estimated Sub-Gradient Solver for SVM.” Proceedings of the 24th International Conference on Machine Learning, ICML ’07, 2007, pp. 807–814.
[4] Xu, Wei. “Towards Optimal One Pass Large Scale Learning with Averaged Stochastic Gradient Descent.” CoRR, abs/1107.2490, 2011.
Version History
Introduced in R2020b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)