fitrgp
Fit a Gaussian process regression (GPR) model
Syntax
Description
Mdl = fitrgp(Tbl,ResponseVarName)Tbl, where ResponseVarName is the name of the response variable in Tbl.
Mdl = fitrgp(___,Name=Value)
For example, you can specify the fitting method, the prediction method, the covariance function, or the active set selection method. You can also train a cross-validated model.
Mdl is a RegressionGP object. For
object functions and properties of this object, see RegressionGP.
If you train a cross-validated model, then Mdl is a
RegressionPartitionedGP object. For further analysis on the
cross-validated object, use the object functions of the RegressionPartitionedGP object.
[
also returns Mdl,AggregateOptimizationResults] = fitrgp(___)AggregateOptimizationResults, which contains
hyperparameter optimization results when you specify the
OptimizeHyperparameters and
HyperparameterOptimizationOptions name-value arguments.
You must also specify the ConstraintType and
ConstraintBounds options of
HyperparameterOptimizationOptions. You can use this
syntax to optimize on compact model size instead of cross-validation loss, and
to perform a set of multiple optimization problems that have the same options
but different constraint bounds.
Examples
This example uses the abalone data [1], [2], from the UCI Machine Learning Repository [3]. Download the data and save it in your current folder with the name
abalone.data.
Store the data into a table. Display the first seven rows.
tbl = readtable('abalone.data','Filetype','text',... 'ReadVariableNames',false); tbl.Properties.VariableNames = {'Sex','Length','Diameter','Height',... 'WWeight','SWeight','VWeight','ShWeight','NoShellRings'}; tbl(1:7,:)
ans =
Sex Length Diameter Height WWeight SWeight VWeight ShWeight NoShellRings
___ ______ ________ ______ _______ _______ _______ ________ ____________
'M' 0.455 0.365 0.095 0.514 0.2245 0.101 0.15 15
'M' 0.35 0.265 0.09 0.2255 0.0995 0.0485 0.07 7
'F' 0.53 0.42 0.135 0.677 0.2565 0.1415 0.21 9
'M' 0.44 0.365 0.125 0.516 0.2155 0.114 0.155 10
'I' 0.33 0.255 0.08 0.205 0.0895 0.0395 0.055 7
'I' 0.425 0.3 0.095 0.3515 0.141 0.0775 0.12 8
'F' 0.53 0.415 0.15 0.7775 0.237 0.1415 0.33 20The dataset has 4177 observations. The goal is to predict the age of abalone from eight physical measurements. The last variable, number of shell rings shows the age of the abalone. The first predictor is a categorical variable. The last variable in the table is the response variable.
Fit a GPR model using the subset of regressors method for parameter estimation and fully independent conditional method for prediction. Standardize the predictors.
gprMdl = fitrgp(tbl,'NoShellRings','KernelFunction','ardsquaredexponential',... 'FitMethod','sr','PredictMethod','fic','Standardize',1)
grMdl =
RegressionGP
PredictorNames: {1x8 cell}
ResponseName: 'Var9'
ResponseTransform: 'none'
NumObservations: 4177
KernelFunction: 'ARDSquaredExponential'
KernelInformation: [1x1 struct]
BasisFunction: 'Constant'
Beta: 10.9148
Sigma: 2.0243
PredictorLocation: [10x1 double]
PredictorScale: [10x1 double]
Alpha: [1000x1 double]
ActiveSetVectors: [1000x10 double]
PredictMethod: 'FIC'
ActiveSetSize: 1000
FitMethod: 'SR'
ActiveSetMethod: 'Random'
IsActiveSetVector: [4177x1 logical]
LogLikelihood: -9.0013e+03
ActiveSetHistory: [1x1 struct]
BCDInformation: []
Predict the responses using the trained model.
ypred = resubPredict(gprMdl);
Plot the true response and the predicted responses.
figure(); plot(tbl.NoShellRings,'r.'); hold on plot(ypred,'b'); xlabel('x'); ylabel('y'); legend({'data','predictions'},'Location','Best'); axis([0 4300 0 30]); hold off;
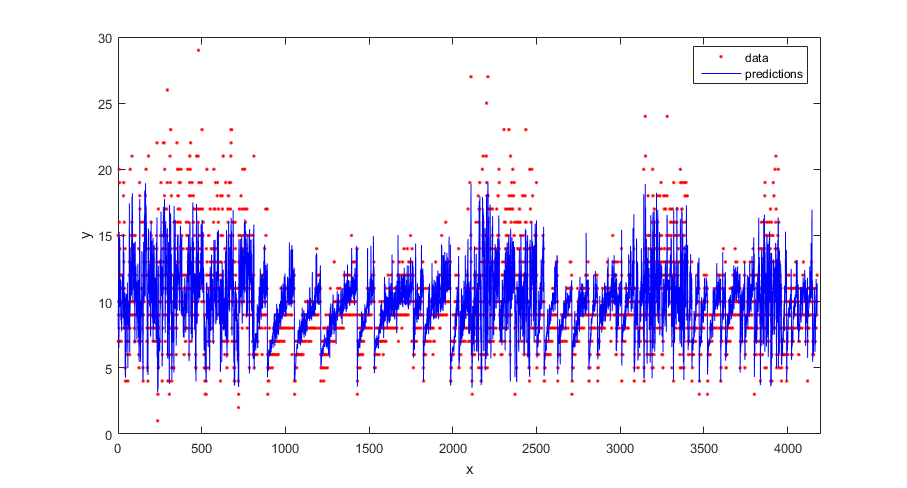
Compute the regression loss on the training data (resubstitution loss) for the trained model.
L = resubLoss(gprMdl)
L =
4.0064Generate sample data.
rng(0,'twister'); % For reproducibility n = 1000; x = linspace(-10,10,n)'; y = 1 + x*5e-2 + sin(x)./x + 0.2*randn(n,1);
Fit a GPR model using a linear basis function and the exact fitting method to estimate the parameters. Also use the exact prediction method.
gprMdl = fitrgp(x,y,'Basis','linear',... 'FitMethod','exact','PredictMethod','exact');
Predict the response corresponding to the rows of x (resubstitution predictions) using the trained model.
ypred = resubPredict(gprMdl);
Plot the true response with the predicted values.
plot(x,y,'b.'); hold on; plot(x,ypred,'r','LineWidth',1.5); xlabel('x'); ylabel('y'); legend('Data','GPR predictions'); hold off
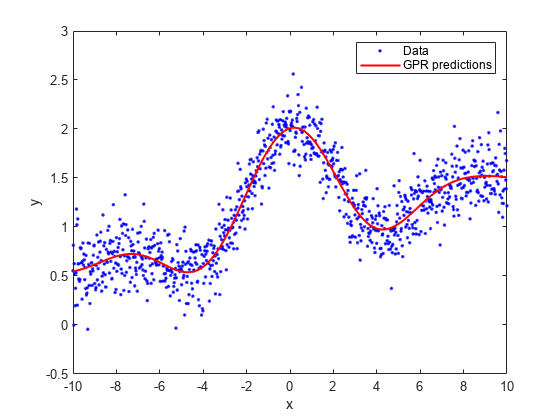
Load the sample data.
load('gprdata2.mat')The data has one predictor variable and continuous response. This is simulated data.
Fit a GPR model using the squared exponential kernel function with default kernel parameters.
gprMdl1 = fitrgp(x,y,'KernelFunction','squaredexponential');
Now, fit a second model, where you specify the initial values for the kernel parameters.
sigma0 = 0.2; kparams0 = [3.5, 6.2]; gprMdl2 = fitrgp(x,y,'KernelFunction','squaredexponential',... 'KernelParameters',kparams0,'Sigma',sigma0);
Compute the resubstitution predictions from both models.
ypred1 = resubPredict(gprMdl1); ypred2 = resubPredict(gprMdl2);
Plot the response predictions from both models and the responses in training data.
figure(); plot(x,y,'r.'); hold on plot(x,ypred1,'b'); plot(x,ypred2,'g'); xlabel('x'); ylabel('y'); legend({'data','default kernel parameters',... 'kparams0 = [3.5,6.2], sigma0 = 0.2'},... 'Location','Best'); title('Impact of initial kernel parameter values'); hold off
![Figure contains an axes object. The axes object with title Impact of initial kernel parameter values, xlabel x, ylabel y contains 3 objects of type line. One or more of the lines displays its values using only markers These objects represent data, default kernel parameters, kparams0 = [3.5,6.2], sigma0 = 0.2.](../examples/stats/win64/ImpactofSpecifyingInitialKernelParameterValuesExample_01.png)
The marginal log likelihood that fitrgp maximizes to estimate GPR parameters has multiple local solutions; the solution that it converges to depends on the initial point. Each local solution corresponds to a particular interpretation of the data. In this example, the solution with the default initial kernel parameters corresponds to a low frequency signal with high noise whereas the second solution with custom initial kernel parameters corresponds to a high frequency signal with low noise.
Load the sample data.
load('gprdata.mat')There are six continuous predictor variables. There are 500 observations in the training data set and 100 observations in the test data set. This is simulated data.
Fit a GPR model using the squared exponential kernel function with a separate length scale for each predictor. This covariance function is defined as:
where represents the length scale for predictor , = 1, 2, ..., and is the signal standard deviation. The unconstrained parametrization is
Initialize length scales of the kernel function at 10 and signal and noise standard deviations at the standard deviation of the response.
sigma0 = std(ytrain); sigmaF0 = sigma0; d = size(Xtrain,2); sigmaM0 = 10*ones(d,1);
Fit the GPR model using the initial kernel parameter values. Standardize the predictors in the training data. Use the exact fitting and prediction methods.
gprMdl = fitrgp(Xtrain,ytrain,'Basis','constant','FitMethod','exact',... 'PredictMethod','exact','KernelFunction','ardsquaredexponential',... 'KernelParameters',[sigmaM0;sigmaF0],'Sigma',sigma0,'Standardize',1);
Compute the regression loss on the test data.
L = loss(gprMdl,Xtest,ytest)
L = 0.6919
Access the kernel information.
gprMdl.KernelInformation
ans = struct with fields:
Name: 'ARDSquaredExponential'
KernelParameters: [7×1 double]
KernelParameterNames: {7×1 cell}
Display the kernel parameter names. When you use an ARD kernel, the length scale parameters correspond to the expanded predictors. Replace the length scale parameter names with the expanded predictor names.
paramNames = gprMdl.KernelInformation.KernelParameterNames
paramNames = 7×1 cell
{'LengthScale1'}
{'LengthScale2'}
{'LengthScale3'}
{'LengthScale4'}
{'LengthScale5'}
{'LengthScale6'}
{'SigmaF' }
paramNames(1:end-1) = gprMdl.ExpandedPredictorNames
paramNames = 7×1 cell
{'x1' }
{'x2' }
{'x3' }
{'x4' }
{'x5' }
{'x6' }
{'SigmaF'}
Display the kernel parameters.
sigmaM = gprMdl.KernelInformation.KernelParameters(1:end-1,1)
sigmaM = 6×1
104 ×
0.0004
0.0007
0.0004
4.7605
0.1018
0.0056
sigmaF = gprMdl.KernelInformation.KernelParameters(end)
sigmaF = 28.1720
sigma = gprMdl.Sigma
sigma = 0.8162
Plot the log of learned length scales.
figure() plot((1:d)',log(sigmaM),'ro-'); xlabel('Length scale number'); ylabel('Log of length scale');
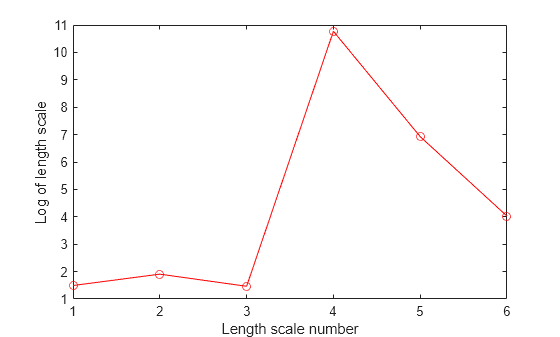
The log of length scale for the 4th and 5th predictor variables are high relative to the others. These predictor variables do not seem to be as influential on the response as the other predictor variables.
Fit the GPR model without using the 4th and 5th variables as the predictor variables.
X = [Xtrain(:,1:3) Xtrain(:,6)]; sigma0 = std(ytrain); sigmaF0 = sigma0; d = size(X,2); sigmaM0 = 10*ones(d,1); gprMdl = fitrgp(X,ytrain,'Basis','constant','FitMethod','exact',... 'PredictMethod','exact','KernelFunction','ardsquaredexponential',... 'KernelParameters',[sigmaM0;sigmaF0],'Sigma',sigma0,'Standardize',1);
Compute the regression error on the test data.
xtest = [Xtest(:,1:3) Xtest(:,6)]; L = loss(gprMdl,xtest,ytest)
L = 0.6928
The loss is similar to the one when all variables are used as predictor variables.
Compute the predicted response for the test data.
ypred = predict(gprMdl,xtest);
Plot the original response along with the fitted values.
figure; plot(ytest,'r'); hold on; plot(ypred,'b'); legend('True response','GPR predicted values','Location','Best'); hold off
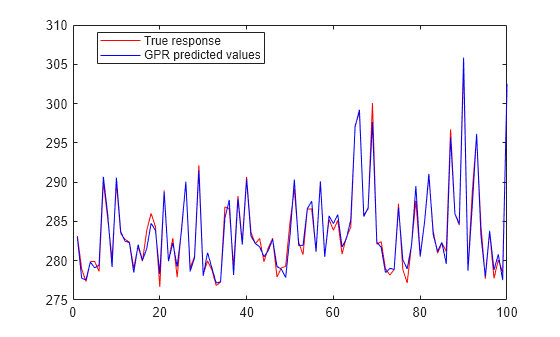
Automatically optimize hyperparameters of a GPR model using fitrgp. Compare the fit of the model with the optimized hyperparameter values to the fit of the model with the default hyperparameter values.
Load the gprdata2 data set.
load gprdata2The simulated data consists of one predictor variable and one continuous response.
Fit a GPR model using the squared exponential kernel function with default kernel parameters.
gprMdl1 = fitrgp(x,y,KernelFunction="squaredexponential");Find hyperparameters that minimize the 5-fold cross-validation loss by using automatic hyperparameter optimization. For reproducibility, set the random seed and use the "expected-improvement-plus" acquisition function.
rng(0,"twister") hpoOptions = hyperparameterOptimizationOptions(AcquisitionFunctionName="expected-improvement-plus"); gprMdl2 = fitrgp(x,y,KernelFunction="squaredexponential", ... OptimizeHyperparameters="auto",HyperparameterOptimizationOptions=hpoOptions);
|=====================================================================================================|
| Iter | Eval | Objective: | Objective | BestSoFar | BestSoFar | Sigma | Standardize |
| | result | log(1+loss) | runtime | (observed) | (estim.) | | |
|=====================================================================================================|
| 1 | Best | 0.42258 | 0.98455 | 0.42258 | 0.42258 | 2.7255 | false |
| 2 | Accept | 1.1091 | 1.2998 | 0.42258 | 0.61465 | 0.0057724 | true |
| 3 | Accept | 1.6498 | 0.63221 | 0.42258 | 0.75989 | 25.905 | true |
| 4 | Best | 0.29824 | 1.0615 | 0.29824 | 0.29834 | 0.0098001 | false |
| 5 | Accept | 1.0387 | 1.3048 | 0.29824 | 0.49293 | 0.00010004 | false |
| 6 | Best | 0.037911 | 1.0524 | 0.037911 | 0.038158 | 0.13791 | false |
| 7 | Accept | 0.042394 | 0.86498 | 0.037911 | 0.038142 | 0.079273 | false |
| 8 | Accept | 0.038899 | 1.1123 | 0.037911 | 0.037126 | 0.11545 | false |
| 9 | Accept | 0.038969 | 0.97929 | 0.037911 | 0.037592 | 0.11307 | false |
| 10 | Accept | 0.039193 | 0.92651 | 0.037911 | 0.037928 | 0.1124 | false |
| 11 | Accept | 1.1496 | 1.1498 | 0.037911 | 0.03793 | 0.0001 | true |
| 12 | Accept | 0.13152 | 0.87078 | 0.037911 | 0.038375 | 0.37155 | true |
| 13 | Accept | 0.037915 | 0.87061 | 0.037911 | 0.038351 | 0.13701 | true |
| 14 | Best | 0.037785 | 0.87043 | 0.037785 | 0.038272 | 0.18141 | true |
| 15 | Best | 0.037697 | 0.8999 | 0.037697 | 0.03876 | 0.32603 | false |
| 16 | Accept | 0.037716 | 1.5285 | 0.037697 | 0.038783 | 0.22469 | false |
| 17 | Accept | 0.037832 | 1.0669 | 0.037697 | 0.036007 | 0.16182 | true |
| 18 | Accept | 0.037702 | 0.90477 | 0.037697 | 0.036011 | 0.23962 | false |
| 19 | Accept | 0.037838 | 1.2147 | 0.037697 | 0.036467 | 0.15992 | true |
| 20 | Best | 0.037694 | 0.92677 | 0.037694 | 0.036469 | 0.24919 | false |
|=====================================================================================================|
| Iter | Eval | Objective: | Objective | BestSoFar | BestSoFar | Sigma | Standardize |
| | result | log(1+loss) | runtime | (observed) | (estim.) | | |
|=====================================================================================================|
| 21 | Accept | 0.037841 | 1.0112 | 0.037694 | 0.03675 | 0.15878 | true |
| 22 | Accept | 0.037765 | 1.005 | 0.037694 | 0.036751 | 0.19128 | false |
| 23 | Best | 0.037683 | 0.92424 | 0.037683 | 0.036752 | 0.27578 | false |
| 24 | Accept | 2.1082 | 0.77623 | 0.037683 | 0.036754 | 29.263 | false |
| 25 | Accept | 0.42164 | 0.67778 | 0.037683 | 0.034063 | 0.5675 | false |
| 26 | Accept | 0.29404 | 0.90962 | 0.037683 | 0.034072 | 0.029617 | false |
| 27 | Accept | 1.0384 | 0.86692 | 0.037683 | 0.034106 | 0.0021049 | false |
| 28 | Accept | 0.28288 | 1.0758 | 0.037683 | 0.034043 | 0.051917 | true |
| 29 | Accept | 0.42298 | 0.79032 | 0.037683 | 0.03404 | 1.3844 | true |
| 30 | Accept | 0.29873 | 1.2886 | 0.037683 | 0.034238 | 0.00073587 | true |
__________________________________________________________
Optimization completed.
MaxObjectiveEvaluations of 30 reached.
Total function evaluations: 30
Total elapsed time: 44.8443 seconds
Total objective function evaluation time: 29.8472
Best observed feasible point:
Sigma Standardize
_______ ___________
0.27578 false
Observed objective function value = 0.037683
Estimated objective function value = 0.034238
Function evaluation time = 0.92424
Best estimated feasible point (according to models):
Sigma Standardize
_______ ___________
0.27578 false
Estimated objective function value = 0.034238
Estimated function evaluation time = 0.95655
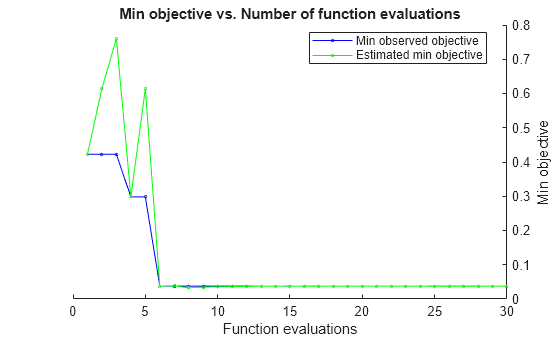
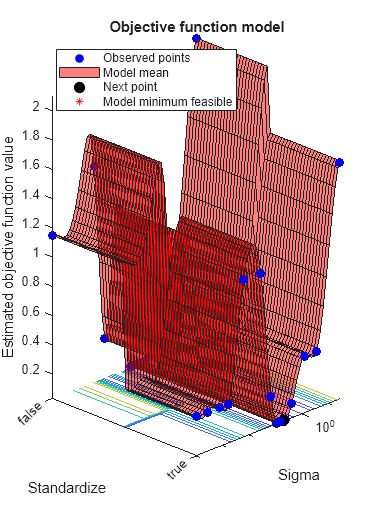
The trained regression model gprMdl2 corresponds to the best estimated feasible point and uses the same hyperparameter values for Sigma and Standardize.
Find the hyperparameter values used to train gprMdl2 by using the bestPoint function. By default, bestPoint uses the same best point criterion used by fitrgp during the hyperparameter optimization ("min-visited-upper-confidence-interval"). In general, fit functions determine the best hyperparameter values based on the "min-visited-upper-confidence-interval" criterion (instead of the "min-observed" criterion) to avoid overfitting to noise in the data set.
bestEstimatedPoint = bestPoint(gprMdl2.HyperparameterOptimizationResults)
bestEstimatedPoint=1×2 table
Sigma Standardize
_______ ___________
0.27578 false
Verify that the results match the properties of gprMdl2. Note that the PredictorLocation and PredictorScale properties of a RegressionGP object are nonempty when the GPR model uses standardization.
modelProperties = table(gprMdl2.Sigma, ... struct(Mean=gprMdl2.PredictorLocation, ... StandardDeviation=gprMdl2.PredictorScale), ... VariableNames=["Sigma","Standardize"])
modelProperties=1×2 table
Sigma Standardize
_______ ___________
0.27578 1×1 struct
modelProperties.Standardize
ans = struct with fields:
Mean: []
StandardDeviation: []
Visually compare the pre- and post-optimization fits.
ypred1 = resubPredict(gprMdl1); ypred2 = resubPredict(gprMdl2); figure plot(x,y,".") hold on plot(x,ypred1) plot(x,ypred2) xlabel("x"); ylabel("y"); legend(["Data","Default fit","Optimized fit"], ... Location="best") title("Impact of Optimization") hold off
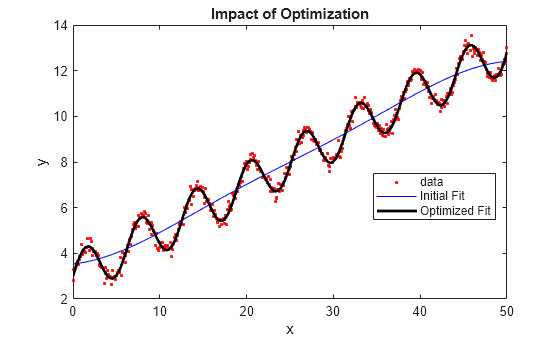
The model with the optimized hyperparameters provides a better fit to the training data.
This example uses the abalone data [1], [2], from the UCI Machine Learning Repository [3]. Download the data and save it in your current folder with the name abalone.data.
Store the data into a table. Display the first seven rows.
tbl = readtable('abalone.data','Filetype','text','ReadVariableNames',false); tbl.Properties.VariableNames = {'Sex','Length','Diameter','Height','WWeight','SWeight','VWeight','ShWeight','NoShellRings'}; tbl(1:7,:)
ans =
Sex Length Diameter Height WWeight SWeight VWeight ShWeight NoShellRings
___ ______ ________ ______ _______ _______ _______ ________ ____________
'M' 0.455 0.365 0.095 0.514 0.2245 0.101 0.15 15
'M' 0.35 0.265 0.09 0.2255 0.0995 0.0485 0.07 7
'F' 0.53 0.42 0.135 0.677 0.2565 0.1415 0.21 9
'M' 0.44 0.365 0.125 0.516 0.2155 0.114 0.155 10
'I' 0.33 0.255 0.08 0.205 0.0895 0.0395 0.055 7
'I' 0.425 0.3 0.095 0.3515 0.141 0.0775 0.12 8
'F' 0.53 0.415 0.15 0.7775 0.237 0.1415 0.33 20The dataset has 4177 observations. The goal is to predict the age of abalone from eight physical measurements. The last variable, number of shell rings shows the age of the abalone. The first predictor is a categorical variable. The last variable in the table is the response variable.
Train a cross-validated GPR model using the 25% of the data for validation.
rng('default') % For reproducibility cvgprMdl = fitrgp(tbl,'NoShellRings','Standardize',1,'Holdout',0.25);
Compute the average loss on folds using models trained on out-of-fold observations.
kfoldLoss(cvgprMdl)
ans = 4.6409
Predict the responses for out-of-fold data.
ypred = kfoldPredict(cvgprMdl);
Plot the true responses used for testing and the predictions.
figure(); plot(ypred(cvgprMdl.Partition.test)); hold on; y = table2array(tbl(:,end)); plot(y(cvgprMdl.Partition.test),'r.'); axis([0 1050 0 30]); xlabel('x') ylabel('y') hold off;
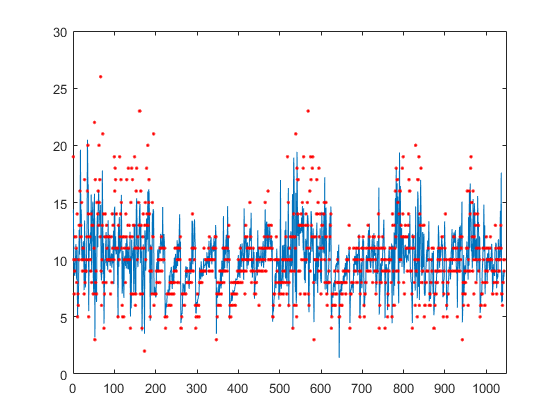
Generate the sample data.
rng(0,'twister'); % For reproducibility n = 1000; x = linspace(-10,10,n)'; y = 1 + x*5e-2 + sin(x)./x + 0.2*randn(n,1);
Define the squared exponential kernel function as a custom kernel function.
You can compute the squared exponential kernel function as
where is the signal standard deviation, is the length scale. Both and must be greater than zero. This condition can be enforced by the unconstrained parameterization, and , for some unconstrained parameterization vector .
Hence, you can define the squared exponential kernel function as a custom kernel function as follows:
kfcn = @(XN,XM,theta) (exp(theta(2))^2)*exp(-(pdist2(XN,XM).^2)/(2*exp(theta(1))^2));
Here pdist2(XN,XM).^2 computes the distance matrix.
Fit a GPR model using the custom kernel function, kfcn. Specify the initial values of the kernel parameters (Because you use a custom kernel function, you must provide initial values for the unconstrained parameterization vector, theta).
theta0 = [1.5,0.2]; gprMdl = fitrgp(x,y,'KernelFunction',kfcn,'KernelParameters',theta0);
fitrgp uses analytical derivatives to estimate parameters when using a built-in kernel function, whereas when using a custom kernel function it uses numerical derivatives.
Compute the resubstitution loss for this model.
L = resubLoss(gprMdl)
L = 0.0391
Fit the GPR model using the built-in squared exponential kernel function option. Specify the initial values of the kernel parameters (Because you use the built-in custom kernel function and specifying initial parameter values, you must provide the initial values for the signal standard deviation and length scale(s) directly).
sigmaL0 = exp(1.5); sigmaF0 = exp(0.2); gprMdl2 = fitrgp(x,y,'KernelFunction','squaredexponential','KernelParameters',[sigmaL0,sigmaF0]);
Compute the resubstitution loss for this model.
L2 = resubLoss(gprMdl2)
L2 = 0.0391
The two loss values are the same as expected.
Train a GPR model on generated data with many predictors. Specify the initial step size for the LBFGS optimizer.
Set the seed and type of the random number generator for reproducibility of the results.
rng(0,'twister'); % For reproducibility
Generate sample data with 300 observations and 3000 predictors, where the response variable depends on the 4th, 7th, and 13th predictors.
N = 300; P = 3000; X = rand(N,P); y = cos(X(:,7)) + sin(X(:,4).*X(:,13)) + 0.1*randn(N,1);
Set initial values for the kernel parameters.
sigmaL0 = sqrt(P)*ones(P,1); % Length scale for predictors sigmaF0 = 1; % Signal standard deviation
Set initial noise standard deviation to 1.
sigmaN0 = 1;
Specify 1e-2 as the termination tolerance for the relative gradient norm.
opts = statset('fitrgp');
opts.TolFun = 1e-2;Fit a GPR model using the initial kernel parameter values, initial noise standard deviation, and an automatic relevance determination (ARD) squared exponential kernel function.
Specify the initial step size as 1 for determining the initial Hessian approximation for an LBFGS optimizer.
gpr = fitrgp(X,y,'KernelFunction','ardsquaredexponential','Verbose',1, ... 'Optimizer','lbfgs','OptimizerOptions',opts, ... 'KernelParameters',[sigmaL0;sigmaF0],'Sigma',sigmaN0,'InitialStepSize',1);
o Parameter estimation: FitMethod = Exact, Optimizer = lbfgs
o Solver = LBFGS, HessianHistorySize = 15, LineSearchMethod = weakwolfe
|====================================================================================================|
| ITER | FUN VALUE | NORM GRAD | NORM STEP | CURV | GAMMA | ALPHA | ACCEPT |
|====================================================================================================|
| 0 | 3.004966e+02 | 2.569e+02 | 0.000e+00 | | 3.893e-03 | 0.000e+00 | YES |
| 1 | 9.525779e+01 | 1.281e+02 | 1.003e+00 | OK | 6.913e-03 | 1.000e+00 | YES |
| 2 | 3.972026e+01 | 1.647e+01 | 7.639e-01 | OK | 4.718e-03 | 5.000e-01 | YES |
| 3 | 3.893873e+01 | 1.073e+01 | 1.057e-01 | OK | 3.243e-03 | 1.000e+00 | YES |
| 4 | 3.859904e+01 | 5.659e+00 | 3.282e-02 | OK | 3.346e-03 | 1.000e+00 | YES |
| 5 | 3.748912e+01 | 1.030e+01 | 1.395e-01 | OK | 1.460e-03 | 1.000e+00 | YES |
| 6 | 2.028104e+01 | 1.380e+02 | 2.010e+00 | OK | 2.326e-03 | 1.000e+00 | YES |
| 7 | 2.001849e+01 | 1.510e+01 | 9.685e-01 | OK | 2.344e-03 | 1.000e+00 | YES |
| 8 | -7.706109e+00 | 8.340e+01 | 1.125e+00 | OK | 5.771e-04 | 1.000e+00 | YES |
| 9 | -1.786074e+01 | 2.323e+02 | 2.647e+00 | OK | 4.217e-03 | 1.250e-01 | YES |
| 10 | -4.058422e+01 | 1.972e+02 | 6.796e-01 | OK | 7.035e-03 | 1.000e+00 | YES |
| 11 | -7.850209e+01 | 4.432e+01 | 8.335e-01 | OK | 3.099e-03 | 1.000e+00 | YES |
| 12 | -1.312162e+02 | 3.334e+01 | 1.277e+00 | OK | 5.432e-02 | 1.000e+00 | YES |
| 13 | -2.005064e+02 | 9.519e+01 | 2.828e+00 | OK | 5.292e-03 | 1.000e+00 | YES |
| 14 | -2.070150e+02 | 1.898e+01 | 1.641e+00 | OK | 6.817e-03 | 1.000e+00 | YES |
| 15 | -2.108086e+02 | 3.793e+01 | 7.685e-01 | OK | 3.479e-03 | 1.000e+00 | YES |
| 16 | -2.122920e+02 | 7.057e+00 | 1.591e-01 | OK | 2.055e-03 | 1.000e+00 | YES |
| 17 | -2.125610e+02 | 4.337e+00 | 4.818e-02 | OK | 1.974e-03 | 1.000e+00 | YES |
| 18 | -2.130162e+02 | 1.178e+01 | 8.891e-02 | OK | 2.856e-03 | 1.000e+00 | YES |
| 19 | -2.139378e+02 | 1.933e+01 | 2.371e-01 | OK | 1.029e-02 | 1.000e+00 | YES |
|====================================================================================================|
| ITER | FUN VALUE | NORM GRAD | NORM STEP | CURV | GAMMA | ALPHA | ACCEPT |
|====================================================================================================|
| 20 | -2.151111e+02 | 1.550e+01 | 3.015e-01 | OK | 2.765e-02 | 1.000e+00 | YES |
| 21 | -2.173046e+02 | 5.856e+00 | 6.537e-01 | OK | 1.414e-02 | 1.000e+00 | YES |
| 22 | -2.201781e+02 | 8.918e+00 | 8.484e-01 | OK | 6.381e-03 | 1.000e+00 | YES |
| 23 | -2.288858e+02 | 4.846e+01 | 2.311e+00 | OK | 2.661e-03 | 1.000e+00 | YES |
| 24 | -2.392171e+02 | 1.190e+02 | 6.283e+00 | OK | 8.113e-03 | 1.000e+00 | YES |
| 25 | -2.511145e+02 | 1.008e+02 | 1.198e+00 | OK | 1.605e-02 | 1.000e+00 | YES |
| 26 | -2.742547e+02 | 2.207e+01 | 1.231e+00 | OK | 3.191e-03 | 1.000e+00 | YES |
| 27 | -2.849931e+02 | 5.067e+01 | 3.660e+00 | OK | 5.184e-03 | 1.000e+00 | YES |
| 28 | -2.899797e+02 | 2.068e+01 | 1.162e+00 | OK | 6.270e-03 | 1.000e+00 | YES |
| 29 | -2.916723e+02 | 1.816e+01 | 3.213e-01 | OK | 1.415e-02 | 1.000e+00 | YES |
| 30 | -2.947674e+02 | 6.965e+00 | 1.126e+00 | OK | 6.339e-03 | 1.000e+00 | YES |
| 31 | -2.962491e+02 | 1.349e+01 | 2.352e-01 | OK | 8.999e-03 | 1.000e+00 | YES |
| 32 | -3.004921e+02 | 1.586e+01 | 9.880e-01 | OK | 3.940e-02 | 1.000e+00 | YES |
| 33 | -3.118906e+02 | 1.889e+01 | 3.318e+00 | OK | 1.213e-01 | 1.000e+00 | YES |
| 34 | -3.189215e+02 | 7.086e+01 | 3.070e+00 | OK | 8.095e-03 | 1.000e+00 | YES |
| 35 | -3.245557e+02 | 4.366e+00 | 1.397e+00 | OK | 2.718e-03 | 1.000e+00 | YES |
| 36 | -3.254613e+02 | 3.751e+00 | 6.546e-01 | OK | 1.004e-02 | 1.000e+00 | YES |
| 37 | -3.262823e+02 | 4.011e+00 | 2.026e-01 | OK | 2.441e-02 | 1.000e+00 | YES |
| 38 | -3.325606e+02 | 1.773e+01 | 2.427e+00 | OK | 5.234e-02 | 1.000e+00 | YES |
| 39 | -3.350374e+02 | 1.201e+01 | 1.603e+00 | OK | 2.674e-02 | 1.000e+00 | YES |
|====================================================================================================|
| ITER | FUN VALUE | NORM GRAD | NORM STEP | CURV | GAMMA | ALPHA | ACCEPT |
|====================================================================================================|
| 40 | -3.379112e+02 | 5.280e+00 | 1.393e+00 | OK | 1.177e-02 | 1.000e+00 | YES |
| 41 | -3.389136e+02 | 3.061e+00 | 7.121e-01 | OK | 2.935e-02 | 1.000e+00 | YES |
| 42 | -3.401070e+02 | 4.094e+00 | 6.224e-01 | OK | 3.399e-02 | 1.000e+00 | YES |
| 43 | -3.436291e+02 | 8.833e+00 | 1.707e+00 | OK | 5.231e-02 | 1.000e+00 | YES |
| 44 | -3.456295e+02 | 5.891e+00 | 1.424e+00 | OK | 3.772e-02 | 1.000e+00 | YES |
| 45 | -3.460069e+02 | 1.126e+01 | 2.580e+00 | OK | 3.907e-02 | 1.000e+00 | YES |
| 46 | -3.481756e+02 | 1.546e+00 | 8.142e-01 | OK | 1.565e-02 | 1.000e+00 | YES |
Infinity norm of the final gradient = 1.546e+00
Two norm of the final step = 8.142e-01, TolX = 1.000e-12
Relative infinity norm of the final gradient = 6.016e-03, TolFun = 1.000e-02
EXIT: Local minimum found.
o Alpha estimation: PredictMethod = Exact
Because the GPR model uses an ARD kernel with many predictors, using an LBFGS approximation to the Hessian is more memory efficient than storing the full Hessian matrix. Also, using the initial step size to determine the initial Hessian approximation may help speed up optimization.
Find the predictor weights by taking the exponential of the negative learned length scales. Normalize the weights.
sigmaL = gpr.KernelInformation.KernelParameters(1:end-1); % Learned length scales weights = exp(-sigmaL); % Predictor weights weights = weights/sum(weights); % Normalized predictor weights
Plot the normalized predictor weights.
figure; semilogx(weights,'ro'); xlabel('Predictor index'); ylabel('Predictor weight');
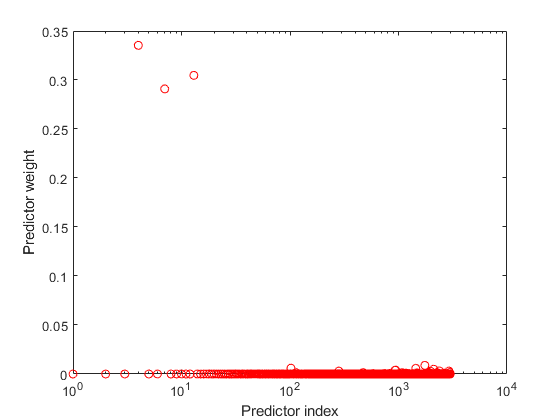
The trained GPR model assigns the largest weights to the 4th, 7th, and 13th predictors. The irrelevant predictors have weights close to zero.
Input Arguments
Sample data used to train the model, specified as a table. Each row of Tbl corresponds to one observation, and each column corresponds to one variable. Tbl contains the predictor variables, and optionally it can also contain one column for the response variable. Multicolumn variables and cell arrays other than cell arrays of character vectors are not allowed.
If
Tblcontains the response variable, and you want to use all the remaining variables as predictors, then specify the response variable usingResponseVarName.If
Tblcontains the response variable, and you want to use only a subset of the predictors in training the model, then specify the response variable and the predictor variables usingformula.If
Tbldoes not contain the response variable, then specify a response variable usingy. The length of the response variable and the number of rows inTblmust be equal.
For more information on the table data type, see table.
If your predictor data contains categorical variables, then fitrgp creates dummy variables. For details, see CategoricalPredictors.
Data Types: table
Response variable name, specified as the name of a variable in
Tbl. You must specify
ResponseVarName as a character vector or string
scalar. For example, if the response variable y is stored
in Tbl (as Tbl.y), then specify it
as "y". Otherwise, the software treats all the columns of
Tbl, including y, as predictors
when training the model.
Data Types: char | string
Response and predictor variables to use in model training, specified as a
character vector or string scalar in the form of
"y~x1+x2+x3". In this form, y
represents the response variable; x1,
x2, x3 represent the predictor
variables to use in training the model.
Use a formula if you want to specify a subset of variables in Tbl as predictors to use when training the model. If you specify a formula, then any variables that do not appear in formula are not used to train the model.
The variable names in the formula must be both variable names in Tbl
(Tbl.Properties.VariableNames) and valid MATLAB® identifiers. You can verify the variable names in Tbl by
using the isvarname function. If the variable names
are not valid, then you can convert them by using the matlab.lang.makeValidName function.
The formula does not indicate the form of the BasisFunction.
Example: "PetalLength~PetalWidth+Species" identifies the
variable PetalLength as the response variable, and
PetalWidth and Species as the
predictor variables.
Data Types: char | string
Predictor data for the GPR model, specified as an n-by-d matrix. n is the number of observations (rows), and d is the number of predictors (columns).
The length of y and the number of rows of X must be equal.
To specify the names of the predictors in the order of their appearance in
X, use the PredictorNames
name-value argument.
Data Types: double
Response data for the GPR model, specified as an n-by-1 vector. You can omit y if you provide the Tbl training data that also includes y. In that case, use ResponseVarName to identify the response variable or use formula to identify the response and predictor variables.
Data Types: double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: FitMethod="sr",BasisFunction="linear",ActiveSetMethod="sgma",PredictMethod="fic"
trains the GPR model using the subset of regressors approximation method for
parameter estimation, uses a linear basis function, uses sparse greedy matrix
approximation for active selection, and fully independent conditional approximation
method for prediction.
Note
You cannot use any cross-validation name-value argument together with the
OptimizeHyperparameters name-value argument. You can modify the
cross-validation for OptimizeHyperparameters only by using the
HyperparameterOptimizationOptions name-value argument.
Fitting
Method to estimate the parameters of the GPR model, specified as one of the following.
| Fit Method | Description |
|---|---|
"none" | No estimation. Use the initial parameter values as the known parameter values. |
"exact" | Exact Gaussian process regression. This value is the default if n ≤ 2000, where n is the number of observations. |
"sd" | Subset of data points approximation. This value is the default if
n > 2000, where n is the
number of observations. "sd" is a sparse
method. |
"sr" | Subset of regressors approximation. "sr" is a
sparse method. |
"fic" | Fully independent conditional approximation. "fic"
is a sparse method. |
Example: FitMethod="fic"
Explicit basis in the GPR model, specified as "constant",
"none", "linear",
"pureQuadratic", or a function handle. If n is
the number of observations, the basis function adds the term H*β to the model, where H is the basis matrix and β is a p-by-1 vector of basis
coefficients.
| Explicit Basis | Basis Matrix |
|---|---|
"none" | Empty matrix |
"constant" |
H is an n-by-1 vector of 1s, where n is the number of observations. |
"linear" |
X is the expanded predictor data after
the software creates dummy variables for the categorical variables.
For details about creating dummy variables, see
|
"pureQuadratic" |
where
For this basis option, the software does not support X with categorical predictors. |
| Function handle | Function handle where X is an n-by-d matrix of predictors, d is the number of predictors after the software creates dummy variables for the categorical variables, and H is an n-by-p matrix of basis functions. |
Example: BasisFunction="pureQuadratic"
Data Types: char | string | function_handle
Initial value of the coefficients for the explicit basis, specified as a p-by-1 vector, where p is the number of columns in the basis matrix H.
The basis matrix depends on the specified basis function. For more
information, see BasisFunction.
The training function uses the coefficient initial values as the known
coefficient values only when FitMethod is
"none".
Data Types: double
Initial value for the noise standard deviation of the Gaussian process model, specified as a positive scalar value.
The training function parameterizes the noise standard deviation as the sum of
SigmaLowerBound and
exp(η), where η is an
unconstrained value. Therefore, Sigma must be larger than
SigmaLowerBound by a small tolerance so that the function can
initialize η to a finite value. Otherwise, the function resets
Sigma to a compatible value.
The tolerance is 1e-3 when ConstantSigma is
false (default) and 1e-6 otherwise. If the
tolerance is not small enough relative to the scale of the response variable, you can
scale up the response variable so that the tolerance value can be considered small for
the response variable.
Example: Sigma=2
Data Types: double
Constant value of Sigma for the noise standard deviation of the
Gaussian process model, specified as a numeric or logical 0
(false) or 1 (true). When
ConstantSigma is true, the training function
does not optimize the value of Sigma, but instead uses the initial
value throughout its computations.
Example: ConstantSigma=true
Data Types: logical
Lower bound on the noise standard deviation
(Sigma), specified as a
positive scalar value.
Sigma must be larger than
SigmaLowerBound by a small
tolerance.
Example: SigmaLowerBound=0.02
Data Types: double
Categorical predictors list, specified as one of the values in this table.
| Value | Description |
|---|---|
| Vector of positive integers |
Each entry in the vector is an index value indicating that the corresponding predictor is
categorical. The index values are between 1 and If |
| Logical vector |
A |
| Character matrix | Each row of the matrix is the name of a predictor variable. The names must match the entries in PredictorNames. Pad the names with extra blanks so each row of the character matrix has the same length. |
| String array or cell array of character vectors | Each element in the array is the name of a predictor variable. The names must match the entries in PredictorNames. |
"all" | All predictors are categorical. |
By default, if the
predictor data is in a table (Tbl), fitrgp
assumes that a variable is categorical if it is a logical vector, categorical vector, character
array, string array, or cell array of character vectors. If the predictor data is a matrix
(X), fitrgp assumes that all predictors are
continuous. To identify any other predictors as categorical predictors, specify them by using
the CategoricalPredictors name-value argument.
For the identified categorical predictors, fitrgp creates dummy variables using two different schemes, depending on whether a categorical variable is unordered or ordered. For an unordered categorical variable, fitrgp creates one dummy variable for each level of the categorical variable. For an ordered categorical variable, fitrgp creates one less dummy variable than the number of categories. For details, see Automatic Creation of Dummy Variables.
Example: CategoricalPredictors="all"
Data Types: single | double | logical | char | string | cell
Indicator to standardize data, specified as a numeric or logical 0
(false) or 1 (true).
If you set Standardize=1, then the software centers and scales each column
of the predictor data by the column mean and standard deviation. The software does not
standardize the data contained in the dummy variable columns generated for categorical
predictors.
Example: Standardize=1
Example: Standardize=true
Data Types: logical
Regularization standard deviation for the subset of regressors ("sr") and
fully independent conditional
("fic") approximation methods,
specified as a positive scalar value. For more
information, see
FitMethod.
Example: Regularization=0.2
Data Types: double
Method for computing the loglikelihood and gradient for parameter estimation,
specified as "qr" or "v". This argument is valid
when FitMethod is "sr" or
"fic".
"qr"— Use the QR-factorization-based approach, which provides better accuracy."v"— Use the V-method-based approach, which provides faster computation.
For more information about these approaches, see Foster, et. al. [7].
Example: ComputationMethod="v"
Kernel (Covariance) Function
Form of the covariance function, specified as one of the following.
| Value | Description |
|---|---|
"exponential" | Exponential kernel |
"squaredexponential" | Squared exponential kernel |
"matern32" | Matern kernel with parameter 3/2 |
"matern52" | Matern kernel with parameter 5/2 |
"rationalquadratic" | Rational quadratic kernel |
"ardexponential" | Exponential kernel with a separate length scale per predictor |
"ardsquaredexponential" | Squared exponential kernel with a separate length scale per predictor |
"ardmatern32" | Matern kernel with parameter 3/2 and a separate length scale per predictor |
"ardmatern52" | Matern kernel with parameter 5/2 and a separate length scale per predictor |
"ardrationalquadratic" | Rational quadratic kernel with a separate length scale per predictor |
| Function handle | Function handle in the form:Kmn =
kfcn(Xm,Xn,theta), where Xm is an
m-by-d matrix,
Xn is an
n-by-d matrix, and
Kmn is an
m-by-n matrix of kernel
products such that
Kmn(i,j) is
the kernel product between Xm(i,:)
and Xn(j,:). d
is the number of predictor variables after the software creates dummy
variables for the categorical variables. For details about creating
dummy variables, see
CategoricalPredictors.theta
is the r-by-1 unconstrained parameter vector for
kfcn. |
For more information on the kernel functions, see Kernel (Covariance) Function Options.
Example: KernelFunction="matern32"
Data Types: char | string | function_handle
Initial values for the kernel parameters, specified as a numeric vector. The size of
the vector and the values depend on the form of the covariance function, specified by
the KernelFunction name-value argument.
KernelFunction Value | KernelParameters Value |
|---|---|
"exponential",
"squaredexponential",
"matern32", or
"matern52" | 2-by-1 vector phi, where phi(1)
contains the length scale and phi(2) contains the
signal standard deviation. The default initial value of the length scale parameter is the mean of the standard deviations of the predictors. The signal standard deviation is the standard deviation of the responses divided by the square root of 2. That is, phi =
[mean(std(X));std(y)/sqrt(2)]. |
"rationalquadratic" | 3-by-1 vector phi, where phi(1)
contains the length scale, phi(2) contains the
scale-mixture parameter, and phi(3) contains the
signal standard deviation. The default initial value of the length scale parameter is the mean of the standard deviations of the predictors. The signal standard deviation is the standard deviation of the responses divided by the square root of 2. The default initial value for the scale-mixture parameter is 1. That is, phi =
[mean(std(X));1;std(y)/sqrt(2)]. |
"ardexponential",
"ardsquaredexponential",
"ardmatern32", or
"ardmatern52" | (d+1)-by-1 vector phi, where
phi(i) contains the length scale for predictor
i, and
phi(d+1) contains the signal
standard deviation. d is the number of predictor
variables after the software creates dummy variables for the categorical
variables. For details about creating dummy variables, see
CategoricalPredictors. The default initial values of the length scale parameters are the standard deviations of the predictors. The signal standard deviation is the standard deviation of the responses divided by the square root of 2. That is, phi =
[std(X)';std(y)/sqrt(2)]. |
"ardrationalquadratic" | (d+2)-by-1 vector phi, where
phi(i) contains the length scale for predictor
i, phi(d+1) contains the
scale-mixture parameter, and phi(d+2) contains the
signal standard deviation. d is the number of
predictor variables after the software creates dummy variables for the
categorical variables. For details about creating dummy variables, see
CategoricalPredictors.The default initial values of the length scale parameters are the standard deviations of the predictors. The signal standard deviation is the standard deviation of the responses divided by the square root of 2. The default initial value of the scale-mixture parameter is 1. That is, phi =
[std(X)';1;std(y)/sqrt(2)]. |
| Function handle | r-by-1 vector for the initial value of the
unconstrained parameter vector phi for the custom
kernel function kfcn. When KernelFunction is a function handle, you must
supply initial values for the kernel parameters. |
For more information on the kernel functions, see Kernel (Covariance) Function Options.
Example: KernelParameters=phi
Data Types: double | single
Method for computing inter-point distances to evaluate built-in kernel functions, specified as
"fast" or "accurate". When you specify
"fast", the training function computes as . When you specify "accurate", the training function
computes .
Example: DistanceMethod="accurate"
Active Set Selection
Observations in the active set, specified as an
m-by-1 vector of integers ranging from 1 to
n (m ≤ n)
or a logical vector of length n with at least one
true element. n is the total
number of observations in the training data.
fitrgp uses the observations indicated by ActiveSet to train the GPR model. The active set cannot have duplicate elements.
If you supply ActiveSet, then:
fitrgpdoes not useActiveSetSizeandActiveSetMethod.You cannot perform cross-validation on this model.
Data Types: double | logical
Size of the active set, specified as an integer m, 1 ≤
m ≤ n, where n is the
number of observations. This argument is valid when FitMethod is
"sd", "sr", or
"fic".
The default value is min(1000,n) when
FitMethod is "sr" or
"fic", and min(2000,n) otherwise.
Example: ActiveSetSize=100
Data Types: double
Active set selection method, specified as one of the following values.
| Value | Description |
|---|---|
"random" | Random selection |
"sgma" | Sparse greedy matrix approximation |
"entropy" | Differential entropy-based selection |
"likelihood" | Subset of regressors loglikelihood-based selection |
All active set selection methods (except "random") require the storage of
an n-by-m matrix, where m is
the size of the active set and n is the number of
observations.
Example: ActiveSetMethod="entropy"
Random search set size per greedy inclusion for active set selection, specified as an integer value.
Example: RandomSearchSetSize=30
Data Types: double
Relative tolerance for terminating active set selection, specified as a positive scalar.
Example: ToleranceActiveset=0.0002
Data Types: double
Number of repetitions for interleaved active set selection
and parameter estimation when ActiveSetMethod is not
"random", specified as an integer value.
Example: NumActiveSetRepeats=5
Data Types: double
Prediction
Method used to make predictions from a Gaussian process model given the parameters, specified as one of the following values.
| Value | Description |
|---|---|
"exact" | Exact Gaussian process regression method. This value is the default if n ≤ 10,000. |
"bcd" | Block coordinate descent (BCD). This value is the default if n > 10,000. |
"sd" | Subset of data points approximation |
"sr" | Subset of regressors approximation |
"fic" | Fully independent conditional approximation |
Example: PredictMethod="bcd"
Block size for the block coordinate descent method
("bcd"), specified as an integer
in the range 1 to n, where
n is the number of
observations.
Example: BlockSizeBCD=1500
Data Types: double
Number of greedy selections for the block coordinate descent method
("bcd"), specified as an integer
in the range 1 to BlockSizeBCD.
Example: NumGreedyBCD=150
Data Types: double
Relative tolerance on the gradient norm for terminating the block coordinate descent method
("bcd") iterations, specified
as a positive scalar.
Example: ToleranceBCD=0.002
Data Types: double
Absolute tolerance on the step size for terminating the block coordinate descent method
("bcd") iterations, specified as a positive scalar.
Example: StepToleranceBCD=0.002
Data Types: double
Maximum number of block coordinate descent method ("bcd") iterations,
specified as a positive integer.
Example: IterationLimitBCD=10000
Data Types: double
Optimization
Optimizer to use for parameter estimation, specified as one of the values in this table.
| Value | Description |
|---|---|
"quasinewton" | Dense, symmetric rank-1-based, quasi-Newton approximation to the Hessian |
"lbfgs" | LBFGS-based quasi-Newton approximation to the Hessian |
"fminsearch" | Unconstrained nonlinear optimization using the simplex search method of Lagarias et al. [5] |
"fminunc" | Unconstrained nonlinear optimization (requires an Optimization Toolbox™ license) |
"fmincon" | Constrained nonlinear optimization (requires an Optimization Toolbox license) |
For more information on the optimizers, see Algorithms.
Example: Optimizer="fmincon"
Options for the optimizer set by the Optimizer name-value
argument, specified as a structure or object created by optimset,
statset("fitrgp"), or optimoptions.
| Optimizer | Function for Creating Optimizer Options |
|---|---|
"fminsearch" | optimset (structure) |
"quasinewton" or
"lbfgs" | statset("fitrgp") (structure) |
"fminunc" or "fmincon" | optimoptions (object) |
The default options depend on the specified optimizer.
Example: OptimizerOptions=opt
Initial step size, specified as a real positive scalar or "auto".
InitialStepSize is the approximate maximum absolute value of the first
optimization step when the optimizer is "quasinewton" or
"lbfgs". The initial step size can determine the initial Hessian
approximation during optimization.
By default, the training function does not use the initial step size to determine the initial
Hessian approximation. To use the initial step size, set a value for the
InitialStepSize name-value argument, or specify
InitialStepSize="auto" to have the software determine a value
automatically. For more information on "auto", see Algorithms.
Example: InitialStepSize="auto"
Cross-Validation
Flag to train a cross-validated model, specified as
"on" or "off".
If you specify "on", then the software trains a
cross-validated model with 10 folds.
You can override this cross-validation setting using the
CVPartition, Holdout,
KFold, or Leaveout
name-value argument. You can use only one cross-validation name-value
argument at a time to create a cross-validated model.
Alternatively, cross-validate later by passing
Mdl to crossval.
Example: Crossval="on"
Data Types: char | string
Cross-validation partition, specified as a cvpartition object that specifies the type of cross-validation and the
indexing for the training and validation sets.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: Suppose you create a random partition for 5-fold cross-validation on 500
observations by using cvp = cvpartition(500,KFold=5). Then, you can
specify the cross-validation partition by setting
CVPartition=cvp.
Fraction of the data used for holdout validation, specified as a scalar value in the range
(0,1). If you specify Holdout=p, then the software completes these
steps:
Randomly select and reserve
p*100% of the data as validation data, and train the model using the rest of the data.Store the compact trained model in the
Trainedproperty of the cross-validated model.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: Holdout=0.1
Data Types: double | single
Number of folds to use in the cross-validated model, specified as a positive integer value
greater than 1. If you specify KFold=k, then the software completes
these steps:
Randomly partition the data into
ksets.For each set, reserve the set as validation data, and train the model using the other
k– 1 sets.Store the
kcompact trained models in ak-by-1 cell vector in theTrainedproperty of the cross-validated model.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: KFold=5
Data Types: single | double
Leave-one-out cross-validation flag, specified as "on" or
"off". If you specify Leaveout="on", then for
each of the n observations (where n is the number
of observations, excluding missing observations, specified in the
NumObservations property of the model), the software completes
these steps:
Reserve the one observation as validation data, and train the model using the other n – 1 observations.
Store the n compact trained models in an n-by-1 cell vector in the
Trainedproperty of the cross-validated model.
To create a cross-validated model, you can specify only one of these four name-value
arguments: CVPartition, Holdout,
KFold, or Leaveout.
Example: Leaveout="on"
Data Types: char | string
Hyperparameter Optimization
Parameters to optimize, specified as one of the following:
"none"— Do not optimize."auto"— Use["Sigma","Standardize"]."all"— Optimize all eligible parameters, equivalent to["BasisFunction","KernelFunction","KernelScale","Sigma","Standardize"].String array or cell array of eligible parameter names.
Vector of
optimizableVariableobjects, typically the output ofhyperparameters.
The optimization attempts to minimize the cross-validation loss
(error) for fitrgp by varying the parameters. To control the
cross-validation type and other aspects of the optimization, use the
HyperparameterOptimizationOptions name-value argument. When you use
HyperparameterOptimizationOptions, you can use the (compact) model size
instead of the cross-validation loss as the optimization objective by setting the
ConstraintType and ConstraintBounds options.
Note
The values of OptimizeHyperparameters override any values you
specify using other name-value arguments. For example, setting
OptimizeHyperparameters to "auto" causes
fitrgp to optimize hyperparameters corresponding to the
"auto" option and to ignore any specified values for the
hyperparameters.
The eligible parameters for fitrgp are:
BasisFunction—fitrgpsearches among"constant","none","linear", and"pureQuadratic".KernelFunction—fitrgpsearches among"ardexponential","ardmatern32","ardmatern52","ardrationalquadratic","ardsquaredexponential","exponential","matern32","matern52","rationalquadratic", and"squaredexponential".KernelScale—fitrgpuses theKernelParametersargument to specify the value of the kernel scale parameter, which is held constant during fitting. In this case, all input dimensions are constrained to have the sameKernelScalevalue.fitrgpsearches among positive values log-scaled in the range[1e-3,1e3].KernelScalecannot be optimized for any of the ARD kernels.Sigma—fitrgpsearches among positive values log-scaled in the range[1e-4,max(1e-3,10*ResponseStd)], whereResponseStd = std(y).Internally,
fitrgpsets theConstantSigmaname-value argument totrueso the value ofSigmais constant during the fitting.Standardize—fitrgpsearches amongtrueandfalse.
Set nondefault parameters by passing a vector of optimizableVariable objects that have nondefault values. For example,
load fisheriris params = hyperparameters("fitrgp",meas,species); params(1).Range = [1e-4,1e6];
Pass params as the value of OptimizeHyperparameters.
By default, the iterative display appears at the command line,
and plots appear according to the number of hyperparameters in the optimization. For the
optimization and plots, the objective function is log(1 + cross-validation loss). To control the iterative display, set the Verbose option
of the HyperparameterOptimizationOptions name-value argument. To control
the plots, set the ShowPlots option of the
HyperparameterOptimizationOptions name-value argument.
For an example, see Optimize GPR Model.
Example: OptimizeHyperparameters="auto"
Options for optimization, specified as a HyperparameterOptimizationOptions object or a structure. This argument modifies the effect of the OptimizeHyperparameters name-value argument. If you specify HyperparameterOptimizationOptions, you must also specify OptimizeHyperparameters. All the options are optional. However, you must set ConstraintBounds and ConstraintType to return AggregateOptimizationResults. The options that you can set in a structure are the same as those in the HyperparameterOptimizationOptions object.
| Option | Values | Default |
|---|---|---|
Optimizer |
| "bayesopt" |
ConstraintBounds | Constraint bounds for N optimization problems, specified as an N-by-2 numeric matrix or | [] |
ConstraintTarget | Constraint target for the optimization problems, specified as | If you specify ConstraintBounds and ConstraintType, then the default value is "matlab". Otherwise, the default value is []. |
ConstraintType | Constraint type for the optimization problems, specified as | [] |
AcquisitionFunctionName | Type of acquisition function:
Acquisition functions whose names include | "expected-improvement-per-second-plus" |
LossFun | Type of validation loss to optimize, specified as "auto"
or "mse". In the case of fitrgp,
the two options are equivalent, and the software uses the mean squared
error. | "auto" |
MaxObjectiveEvaluations | Maximum number of objective function evaluations. If you specify multiple optimization problems using ConstraintBounds, the value of MaxObjectiveEvaluations applies to each optimization problem individually. | 30 for "bayesopt" and "randomsearch", and the entire grid for "gridsearch" |
MaxTime | Time limit for the optimization, specified as a nonnegative real scalar. The time limit is in seconds, as measured by | Inf |
NumGridDivisions | For Optimizer="gridsearch", the number of values in each dimension. The value can be a vector of positive integers giving the number of values for each dimension, or a scalar that applies to all dimensions. The software ignores this option for categorical variables. | 10 |
ShowPlots | Logical value indicating whether to show plots of the optimization progress. If this option
is true, the software plots the best observed objective
function value against the iteration number. If you use Bayesian optimization
(Optimizer="bayesopt"), the software
also plots the best estimated objective function value. The best observed
objective function values and best estimated objective function values
correspond to the values in the BestSoFar (observed) and
BestSoFar (estim.) columns of the iterative display,
respectively. You can find these values in the properties ObjectiveMinimumTrace and EstimatedObjectiveMinimumTrace of the
SupervisedLearningBayesianOptimization object. If the
problem includes one or two optimization parameters for Bayesian optimization,
then ShowPlots also plots a model of the objective function
against the parameters. | true |
SaveIntermediateResults | Logical value indicating whether to save the optimization results. If this option is
true, the software overwrites a workspace variable named
SupervisedLearningBayesoptResults at each iteration. The
variable is a SupervisedLearningBayesianOptimization object. If you specify
multiple optimization problems using ConstraintBounds, the
workspace variable is an AggregateBayesianOptimization object named
AggregateBayesoptResults. | false |
Verbose | Display level at the command line:
For details, see the | 1 |
UseParallel | Logical value indicating whether to run the Bayesian optimization in parallel, which requires Parallel Computing Toolbox™. Due to the nonreproducibility of parallel timing, parallel Bayesian optimization does not necessarily yield reproducible results. For details, see Parallel Bayesian Optimization. | false |
Repartition | Logical value indicating whether to repartition the cross-validation at every iteration. If this option is A value of | false |
| Specify only one of the following three options. | ||
CVPartition | cvpartition object created by cvpartition | KFold=5 if you do not specify a cross-validation option |
Holdout | Scalar in the range (0,1) representing the holdout fraction | |
KFold | Integer greater than 1 | |
Example: HyperparameterOptimizationOptions=struct(UseParallel=true)
Other
Predictor variable names, specified as a string array of unique names or cell array of unique character vectors. The functionality of PredictorNames depends on the way you supply the training data.
If you supply
XandY, then you can usePredictorNamesto assign names to the predictor variables inX.The order of the names in
PredictorNamesmust correspond to the predictor order inX. Assuming thatXhas the default orientation, with observations in rows and predictors in columns,PredictorNames{1}is the name ofX(:,1),PredictorNames{2}is the name ofX(:,2), and so on. Also,size(X,2)andnumel(PredictorNames)must be equal.By default,
PredictorNamesis{'x1','x2',...}.
If you supply
Tbl, then you can usePredictorNamesto choose which predictor variables to use in training. That is,fitrgpuses only the variables inPredictorNamesas predictors during training.PredictorNamesmust be a subset ofTbl.Properties.VariableNamesand cannot include the name of any response variable.By default,
PredictorNamescontains the names of all predictor variables.A good practice is to specify the predictors for training using either
PredictorNamesorformula, but not both.
Example: PredictorNames=["SepalLength","SepalWidth","PetalLength","PetalWidth"]
Data Types: string | cell
Response variable name, specified as a character vector or string scalar.
If you supply
Y, then you can useResponseNameto specify a name for the response variable.If you supply
ResponseVarNameorformula, then you cannot useResponseName.
Example: ResponseName="response"
Data Types: char | string
Verbosity level, specified as 0 or 1.
0— The training function suppresses diagnostic messages related to active set selection and block coordinate descent, but displays the messages related to parameter estimation, depending on the value ofDisplayinOptimizerOptions.1— The training function displays the iterative diagnostic messages related to parameter estimation, active set selection, and block coordinate descent.
Example: Verbose=1
Cache size in megabytes (MB), specified as a positive scalar. Cache size is the extra memory
available in addition to the memory required for fitting and
active set selection. The training function uses
CacheSize to:
Decide whether inter-point distances are cached when estimating parameters.
Decide how matrix vector products are computed for the block coordinate descent method and for making predictions.
Example: CacheSize=2000
Data Types: double
Output Arguments
Trained Gaussian process regression model, returned as a RegressionGP object, a RegressionPartitionedGP object, or a cell array of model
objects.
If you set any of the name-value arguments
CrossVal,CVPartition,Holdout,KFold, orLeaveout, thenMdlis aRegressionPartitionedGPobject.If you specify
OptimizeHyperparametersand set theConstraintTypeandConstraintBoundsoptions ofHyperparameterOptimizationOptions, thenMdlis an N-by-1 cell array of model objects, where N is equal to the number of rows inConstraintBounds. If none of the optimization problems yields a feasible model, then each cell array value is[].Otherwise,
Mdlis aRegressionGPmodel object.
To reference properties of a model object, use dot notation.
Aggregate optimization results for multiple optimization problems, returned as an AggregateBayesianOptimization object. To return
AggregateOptimizationResults, you must specify
OptimizeHyperparameters and
HyperparameterOptimizationOptions. You must also specify the
ConstraintType and ConstraintBounds
options of HyperparameterOptimizationOptions. For an example that
shows how to produce this output, see Hyperparameter Optimization with Multiple Constraint Bounds.
More About
For subset of data, subset of regressors, or fully independent
conditional approximation fitting methods (FitMethod equal to
"sd", "sr", or "fic"),
if you do not provide the active set (or inducing input set), fitrgp selects the active set and computes the parameter estimates
in a series of iterations.
In the first iteration, the software uses the initial parameter values in vector η0 = [β0,σ0,θ0] to select an active set A1. The software maximizes the GPR marginal loglikelihood or its approximation using η0 as the initial values and A1 to compute the new parameter estimates η1. Next, the software computes the new loglikelihood L1 using η1 and A1.
In the second iteration, the software selects the active set A2 using the parameter values in η1. Then, using η1 as the initial values and A2, the software maximizes the GPR marginal loglikelihood or its approximation and estimates the new parameter values η2. Then, using η2 and A2, the software computes the new loglikelihood value L2.
The following table summarizes the iterations and the computations at each iteration.
| Iteration Number | Active Set | Parameter Vector | Loglikelihood |
|---|---|---|---|
| 1 | A1 | η1 | L1 |
| 2 | A2 | η2 | L2 |
| 3 | A3 | η3 | L3 |
| … | … | … | … |
The software iterates similarly for a specified number of repetitions. You can specify the
number of replications for active set selection using the
NumActiveSetRepeats name-value argument.
Tips
fitrgpaccepts any combination of fitting, prediction, and active set selection methods. In some cases it might not be possible to compute the standard deviations of the predicted responses, hence the prediction intervals. Seepredict. And in some cases, using the exact method might be expensive due to the size of the training data.The
PredictorNamesproperty stores one element for each of the original predictor variable names. For example, if there are three predictors, one of which is a categorical variable with three levels,PredictorNamesis a 1-by-3 cell array of character vectors.The
ExpandedPredictorNamesproperty stores one element for each of the predictor variables, including the dummy variables. For example, if there are three predictors, one of which is a categorical variable with three levels, thenExpandedPredictorNamesis a 1-by-5 cell array of character vectors.Similarly, the
Betaproperty stores one beta coefficient for each predictor, including the dummy variables.The
Xproperty stores the training data as originally input. It does not include the dummy variables.The default approach to initializing the Hessian approximation in
fitrgpcan be slow when you have a GPR model with many kernel parameters, such as when using an ARD kernel with many predictors. In this case, consider specifying"auto"or a value for the initial step size.You can set
Verbose=1for display of iterative diagnostic messages, and begin training a GPR model using an LBFGS or quasi-Newton optimizer with the defaultfitrgpoptimization. If the iterative diagnostic messages are not displayed after a few seconds, it is possible that initialization of the Hessian approximation is taking too long. In this case, consider restarting training and using the initial step size to speed up optimization.After training a model, you can generate C/C++ code that predicts responses for new data. Generating C/C++ code requires MATLAB Coder™. For details, see Introduction to Code Generation for Statistics and Machine Learning Functions..
Algorithms
Fitting a GPR model involves estimating the following model parameters from the data:
Covariance function parameterized in terms of kernel parameters in vector (see Kernel (Covariance) Function Options)
Noise variance
Coefficient vector of fixed-basis functions
The value of the
KernelParametersname-value argument is a vector that consists of initial values for the signal standard deviation and the characteristic length scales . The software uses these values to determine the kernel parameters. Similarly, theSigmaname-value argument contains the initial value for the noise standard deviation .During optimization, the software creates a vector of unconstrained initial parameter values by using the initial values for the noise standard deviation and the kernel parameters.
The software analytically determines the explicit basis coefficients , specified by the
Betaname-value argument, from estimated values of and . Therefore, does not appear in the vector when the software initializes numerical optimization.Note
If you do not specify the estimation of parameters for the GPR model, the software uses the value of the
Betaname-value argument and other initial parameter values as the known GPR parameter values (seeBeta). In all other cases, the value ofBetais optimized analytically from the objective function.The quasi-Newton optimizer uses a trust-region method with a dense, symmetric rank-1-based (SR1), quasi-Newton approximation to the Hessian. The LBFGS optimizer uses a standard line-search method with a limited-memory Broyden-Fletcher-Goldfarb-Shanno (LBFGS) quasi-Newton approximation to the Hessian. See Nocedal and Wright [6].
If you set the
InitialStepSizename-value argument to"auto"the software determines the initial step size by using .is the initial step vector, and is the vector of unconstrained initial parameter values.
During optimization, the software uses the initial step size as follows:
If you specify
Optimizer="quasinewton"with the initial step size, then the initial Hessian approximation is .If you specify
Optimizer="lbfgs"with the initial step size, then the initial inverse-Hessian approximation is .is the initial gradient vector, and is the identity matrix.
References
[1] Nash, W.J., T. L. Sellers, S. R. Talbot, A. J. Cawthorn, and W. B. Ford. "The Population Biology of Abalone (Haliotis species) in Tasmania. I. Blacklip Abalone (H. rubra) from the North Coast and Islands of Bass Strait." Sea Fisheries Division, Technical Report No. 48, 1994.
[2] Waugh, S. "Extending and Benchmarking Cascade-Correlation: Extensions to the Cascade-Correlation Architecture and Benchmarking of Feed-forward Supervised Artificial Neural Networks." University of Tasmania Department of Computer Science thesis, 1995.
[3] Lichman, M. UCI Machine Learning Repository, Irvine, CA: University of California, School of Information and Computer Science, 2013. http://archive.ics.uci.edu/ml.
[4] Rasmussen, C. E. and C. K. I. Williams. Gaussian Processes for Machine Learning. MIT Press. Cambridge, Massachusetts, 2006.
[5] Lagarias, J. C., J. A. Reeds, M. H. Wright, and P. E. Wright. "Convergence Properties of the Nelder-Mead Simplex Method in Low Dimensions." SIAM Journal of Optimization. Vol. 9, Number 1, 1998, pp. 112–147.
[6] Nocedal, J. and S. J. Wright. Numerical Optimization, Second Edition. Springer Series in Operations Research, Springer Verlag, 2006.
Extended Capabilities
To perform parallel hyperparameter optimization, use the UseParallel=true
option in the HyperparameterOptimizationOptions name-value argument in
the call to the fitrgp function.
For more information on parallel hyperparameter optimization, see Parallel Bayesian Optimization.
For general information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
fitrgpfits the model on a GPU if at least one of the following applies:The input argument
Xis agpuArrayobject.The input argument
Yis agpuArrayobject.The input argument
TblcontainsgpuArraypredictor or response variables.
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2015bfitrgp supports GPU array input arguments.
fitrgp defaults to serial hyperparameter optimization when
HyperparameterOptimizationOptions includes
UseParallel=true and the software cannot open a parallel pool.
In previous releases, the software issues an error under these circumstances.
Starting in R2023b, when you specify "auto" as the OptimizeHyperparameters value, fitrgp includes Standardize as an optimizable hyperparameter.
Starting in R2023b, fitrgp optimizes the kernel scale parameter for Gaussian process regression (GPR) models by using the default search range [1e-3,1e3]. That is, when you specify to optimize the GPR hyperparameter KernelScale by using the OptimizeHyperparameters name-value argument, the function searches among positive values log-scaled in the range [1e-3,1e3].
In previous releases, the default search range for the KernelScale hyperparameter was [1e-3*MaxPredictorRange,MaxPredictorRange], where MaxPredictorRange = max(max(X) - min(X)).
Starting in R2022b, a cross-validated Gaussian process regression (GPR) model is a RegressionPartitionedGP object. In previous releases, a cross-validated GPR
model was a RegressionPartitionedModel object.
You can create a RegressionPartitionedGP object in two ways:
Create a cross-validated model from a GPR model object
RegressionGPby using thecrossvalobject function.Create a cross-validated model by using the
fitrgpfunction and specifying one of the name-value argumentsCrossVal,CVPartition,Holdout,KFold, orLeaveout.
Regardless of whether you train a full or cross-validated GPR model first, you cannot specify an ActiveSet value in the call to fitrgp.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)