layerGraph
(Not recommended) Graph of network layers for deep learning
LayerGraph objects are not recommended. Use dlnetwork objects instead. For more
information, see Version History.
Description
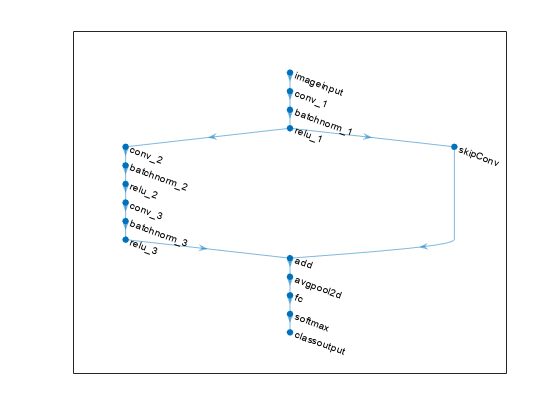
A layer graph specifies the architecture of a neural network as a directed acyclic graph (DAG) of deep learning layers. The layers can have multiple inputs and multiple outputs.
Creation
Description
lgraph = layerGraphaddLayers function.
lgraph = layerGraph(layers)Layers property. The layers in
lgraph are connected in the same sequential order as in
layers.
lgraph = layerGraph(net)SeriesNetwork,
DAGNetwork, or dlnetwork object. For
example, you can extract the layer graph of a pretrained network to perform
transfer learning.
Input Arguments
Properties
Object Functions
addLayers | Add layers to neural network |
removeLayers | Remove layers from neural network |
replaceLayer | Replace layer in neural network |
connectLayers | Connect layers in neural network |
disconnectLayers | Disconnect layers in neural network |
plot | Plot neural network architecture |
Examples
Limitations
Layer graph objects contain no quantization information. Extracting the layer graph from a quantized network and then reassembling the network using
assembleNetworkordlnetworkremoves quantization information from the network.