bayesvarm
Create prior Bayesian vector autoregression (VAR) model object
Syntax
Description
PriorMdl = bayesvarm(numseries,numlags)
The collection of model parameters are in the set
p =
numlagsis the AR polynomial order.The joint prior distribution of (Λ,Σ) is diffuse.
PriorMdl = bayesvarm(numseries,numlags,ModelType=modelType)modelType for Λ and Σ. For this
syntax, modelType can be 'conjugate',
'semiconjugate', 'diffuse', or
'normal'. For example, ModelType="semiconjugate"
specifies semiconjugate priors for the multivariate normal likelihood—specifically, vec(Λ)|Σ
is multivariate normal, Σ is inverse Wishart, and Λ and Σ are independent.
PriorMdl = bayesvarm(numseries,numlags,ModelType=modelType,Name=Value)
Examples
Consider the 3-D VAR(4) model for the US inflation (INFL), unemployment (UNRATE), and federal funds (FEDFUNDS) rates.
For all , is a series of independent 3-D normal innovations with a mean of 0 and variance .
Suppose that the AR coefficient matrices , model constant , and innovations covariance matrix are random variables, and their prior distributions are unknown. In this case, use the noninformative diffuse prior: the joint prior distribution is proportional to .
Create a diffuse prior model for the 3-D VAR(4) model parameters, which is the default prior model type.
numseries = 3; numlags = 4; PriorMdl = bayesvarm(numseries,numlags)
PriorMdl =
diffusebvarm with properties:
Description: "3-Dimensional VAR(4) Model"
NumSeries: 3
P: 4
SeriesNames: ["Y1" "Y2" "Y3"]
IncludeConstant: 1
IncludeTrend: 0
NumPredictors: 0
AR: {[3×3 double] [3×3 double] [3×3 double] [3×3 double]}
Constant: [3×1 double]
Trend: [3×0 double]
Beta: [3×0 double]
Covariance: [3×3 double]
PriorMdl is a diffusevarm Bayesian VAR model object representing the prior distribution of the AR coefficient matrices, model constant vector, and innovations covariance matrix. bayesvarm displays a summary of the prior distributions at the command line.
AR— Prior means of the AR coefficient matrices.Constant— Prior means of the model constant vector.TrendandBeta— Prior means of the linear time trend vector and exogenous regression coefficient matrix, respectively. Because the values are empty arrays, the corresponding parameters are not in the model.Covariance— Prior mean of the innovations covariance matrix.
If you have data, then you can estimate characteristics of the posterior distribution by passing PriorMdl and the data to estimate.
Consider the 3-D VAR(4) model in Default Diffuse Prior Model. Assume the following:
. is a 13-by-3 matrix of prior coefficient means ( is the prior mean matrix of , is the prior mean matrix of ,..., and is the prior mean vector of ). is a 13-by-13 matrix representing the among-coefficient prior covariance matrix within an equation. is the 3-by-3 random innovations covariance matrix.
. is the 3-by-3 scale matrix, and is the degrees of freedom of the inverse Wishart distribution.
The coefficients and the innovations covariance matrix are dependent.
Prior coefficient variances among the equations are proportional.
These assumptions and the data likelihood imply a matrix-normal-inverse-Wishart conjugate model.
Create a matrix-normal-inverse-Wishart conjugate prior model for the VAR model parameters.
numseries = 3;
numlags = 4;
PriorMdl = bayesvarm(numseries,numlags,ModelType="conjugate")PriorMdl =
conjugatebvarm with properties:
Description: "3-Dimensional VAR(4) Model"
NumSeries: 3
P: 4
SeriesNames: ["Y1" "Y2" "Y3"]
IncludeConstant: 1
IncludeTrend: 0
NumPredictors: 0
Mu: [39×1 double]
V: [13×13 double]
Omega: [3×3 double]
DoF: 13
AR: {[3×3 double] [3×3 double] [3×3 double] [3×3 double]}
Constant: [3×1 double]
Trend: [3×0 double]
Beta: [3×0 double]
Covariance: [3×3 double]
PriorMdl is a conjugatebvarm Bayesian VAR model object representing the prior distribution of the coefficients and innovations covariance matrix. bayesvarm displays a summary of the prior distributions at the command line; it returns the prior mean matrix in vectorized form.
The model contains many estimable parameters. To achieve a parsimonious model, bayesvarm applies the Minnesota prior regularization method to the AR coefficients, by default. Inspect the default prior means (centers of shrinkage) of the AR coefficient matrices.
AR1 = PriorMdl.AR{1}AR1 = 3×3
0.5000 0 0
0 0.5000 0
0 0 0.5000
AR2 = PriorMdl.AR{2}AR2 = 3×3
0 0 0
0 0 0
0 0 0
AR3 = PriorMdl.AR{3}AR3 = 3×3
0 0 0
0 0 0
0 0 0
AR4 = PriorMdl.AR{4}AR4 = 3×3
0 0 0
0 0 0
0 0 0
Each series is an AR(1) model with AR coefficient 0.5, a priori.
The tightness on shrinkage of the coefficients is proportional among the equations. Inspect the default tightness values by displaying a heatmap chart of the property V of PriorMdl, which contains a matrix of the scaled tightness on shrinkage of the coefficients for one equation (the unscaled shrinkage is = kron(PriorMdl.Covariance,PriorMdl.V)). Omit the final row and column, which correspond to the model constant.
% Create labels for the chart. numARCoeffMats = PriorMdl.NumSeries*PriorMdl.P; arcoeffnames = strings(numARCoeffMats,1); for r = numlags:-1:1 arcoeffnames(((r-1)*numseries+1):(numseries*r)) = ["\phi_{11,"+r+"}" "\phi_{12,"+r+"}" "\phi_{13,"+r+"}"]; end heatmap(arcoeffnames,arcoeffnames,PriorMdl.V(1:end-1,1:end-1));
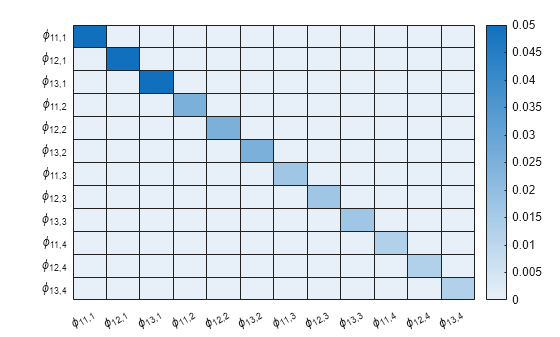
The tightness values decrease with lag, which suggests (a priori) that the means of the corresponding greater-lagged coefficients are more tightly locked around their center of 0.
Display the tightness of the model constant vector.
PriorMdl.V(end,end)
ans = 10000
The center of the model constant vector is 0 but has a large variance, which allows the estimation procedure to defer more to the data than the prior for the posterior mean of the constant vector.
You can specify alternative values after you create a model by using dot notation. For example, increase the tightness of all coefficients by a factor of 100.
PriorMdl.V = 100*PriorMdl.V;
Consider the 3-D VAR(4) model in Default Diffuse Prior Model. Assume these prior distributions, as presented in [1]:
. is a 39-by-1 vector of prior coefficient means (the model has 39 individual coefficients), and is a 39-by-39 prior coefficient covariance matrix.
The innovations covariance is a fixed matrix.
Suppose econometric theory dictates that
Create a normal conjugate prior model for the VAR model coefficients. Specify the value of by using the Sigma name-value argument.
numseries = 3; numlags = 4; Sigma = [10e-5 0 10e-4; 0 0.1 -0.2; 10e-4 -0.2 1.6]; PriorMdl = bayesvarm(numseries,numlags,ModelType="normal", ... Sigma=Sigma)
PriorMdl =
normalbvarm with properties:
Description: "3-Dimensional VAR(4) Model"
NumSeries: 3
P: 4
SeriesNames: ["Y1" "Y2" "Y3"]
IncludeConstant: 1
IncludeTrend: 0
NumPredictors: 0
Mu: [39×1 double]
V: [39×39 double]
Sigma: [3×3 double]
AR: {[3×3 double] [3×3 double] [3×3 double] [3×3 double]}
Constant: [3×1 double]
Trend: [3×0 double]
Beta: [3×0 double]
Covariance: [3×3 double]
PriorMdl is a normalbvarm Bayesian VAR model object representing the prior distribution of the coefficients. Because is fixed for normalbvarm prior models, PriorMdl.Sigma and PriorMdl.Covariance are equal.
PriorMdl.Sigma
ans = 3×3
0.0001 0 0.0010
0 0.1000 -0.2000
0.0010 -0.2000 1.6000
PriorMdl.Covariance
ans = 3×3
0.0001 0 0.0010
0 0.1000 -0.2000
0.0010 -0.2000 1.6000
Consider the 3-D VAR(4) model in Default Diffuse Prior Model. Assume the following:
. is a 39-by-1 vector of prior coefficient means (the model has 39 individual coefficients), and is a 39-by-39 prior coefficient covariance matrix.
. is the 3-by-3 scale matrix, and is the degrees of freedom of the inverse Wishart distribution.
The coefficients and the innovations covariance matrix are independent.
These assumptions and the data likelihood imply a normal-inverse-Wishart semiconjugate model.
The model contains many estimable parameters. To achieve a parsimonious model, bayesvarm enables you to regularize the coefficients by using the Minnesota prior regularization method, rather than specifying each prior mean and variance.
Create a normal-inverse-Wishart semiconjugate prior model for the VAR model parameters. Specify the following:
All series are AR(1) models, a priori, with AR coefficient 0.9. Set the
Centername-value argument to a 3-by-1 vector composed of0.9.The tightness around self lags in is
1. Set theSelfLagname-value argument to1.The tightness around cross lags in is
0.5. Set theCrossLagname-value argument to0.5.All tightness values decay by a factor of the lag degree squared. Set the
Decayname-value argument to2.
numseries = 3; numlags = 4; center = 0.9*ones(numseries,1); PriorMdl = bayesvarm(numseries,numlags,ModelType="semiconjugate", ... Center=center,SelfLag=1,CrossLag=0.5,Decay=2)
PriorMdl =
semiconjugatebvarm with properties:
Description: "3-Dimensional VAR(4) Model"
NumSeries: 3
P: 4
SeriesNames: ["Y1" "Y2" "Y3"]
IncludeConstant: 1
IncludeTrend: 0
NumPredictors: 0
Mu: [39×1 double]
V: [39×39 double]
Omega: [3×3 double]
DoF: 13
AR: {[3×3 double] [3×3 double] [3×3 double] [3×3 double]}
Constant: [3×1 double]
Trend: [3×0 double]
Beta: [3×0 double]
Covariance: [3×3 double]
PriorMdl is a semiconjugatebvarm Bayesian VAR model object representing the prior distribution of the coefficients and innovations covariance matrix. bayesvarm displays a summary of the prior distributions at the command line; it returns the prior mean matrix in vectorized form.
Display the prior means of the AR coefficient matrices.
AR1 = PriorMdl.AR{1}AR1 = 3×3
0.9000 0 0
0 0.9000 0
0 0 0.9000
AR2 = PriorMdl.AR{2}AR2 = 3×3
0 0 0
0 0 0
0 0 0
AR3 = PriorMdl.AR{3}AR3 = 3×3
0 0 0
0 0 0
0 0 0
AR4 = PriorMdl.AR{4}AR4 = 3×3
0 0 0
0 0 0
0 0 0
Each series is an AR(1) model, a priori.
The property V of PriorMdl contains a matrix of the tightness on shrinkage of the coefficients. The rows and columns of V correspond to the elements of the Mu property of PriorMdl.
Elements 1 through 3 correspond to the lag 1 AR coefficients in the first equation ordered by response variable, that is, , , and .
Elements 4 through 6 correspond to the lag 2 AR coefficients in the first equation.
Elements 7 through 9 correspond to the lag 3 AR coefficients in the first equation.
Elements 10 through 12 correspond to the lag 4 AR coefficients in the first equation.
Element 13 is the model constant in the first equation.
MATLAB® repeats the pattern for each equation.
In this example, the tightness of shrinkage is the same for all equations. Display a heatmap chart of the property V of PriorMdl for the tightness values of the AR coefficients in the first equation.
% Create labels for the chart. numARCoeffMats = PriorMdl.NumSeries*PriorMdl.P; arcoeffnames = strings(numARCoeffMats,1); for r = numlags:-1:1 arcoeffnames(((r-1)*numseries+1):(numseries*r)) = ["\phi_{"+r+",11}" "\phi_{"+r+",12}" "\phi_{"+r+"13}"]; end heatmap(arcoeffnames,arcoeffnames,PriorMdl.V(1:numARCoeffMats,1:numARCoeffMats));
![]()
The tightness values decrease with lag, which suggests (a priori) that the means of the corresponding greater-lagged coefficients are more tightly locked around their center of 0. By default, AR coefficients are uncorrelated.
Display the tightness of the model constant vector.
PriorMdl.V(numARCoeffMats + 1,numARCoeffMats + 1)
ans = 10000
The center of the model constant vector is 0 but has a large variance, which allows the estimation procedure to defer more to the data than the prior for the posterior mean of the constant vector.
You can specify alternative values after you create a model by using dot notation. For example, increase the tightness of all coefficients by a factor of 100.
PriorMdl.V = 100*PriorMdl.V;
Input Arguments
Number of time series m, specified as a positive integer. numseries specifies the dimensionality of the multivariate response variable yt and innovation εt.
Data Types: double
Number of lagged responses p to include in the VAR model, specified as a nonnegative integer. bayesvarm includes lags 1 through numlags.
Data Types: double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: IncludeTrend=true,NumPredictors=3 specifies a linear trend
term and a linear regression term for three exogenous variables in all response
equations.
Model Options
Joint prior distribution of (Λ,Σ), specified as a value in the following table. In the table:
λ = vec(Λ).
d =
IncludeConstant+IncludeTrend+NumPredictors.The inverse Wishart hyperparameters Ω and ν correspond to the name-value arguments and output model properties
OmegaandDoF, respectively. You can adjust their values by specifying the name-value arguments or by using dot notation afterbayesvarmreturnsPriorMdl.
| Value | Description |
|---|---|
'conjugate' | Matrix-normal-inverse-Wishart conjugate model. The priors are where Λ and Σ are dependent. |
'semiconjugate' | Normal-inverse-Wishart semiconjugate model. The priors are where Λ and Σ are independent. |
'diffuse' | Diffuse prior distributions. The joint prior pdf is Regularization options do not apply to diffuse priors. |
'normal' | Normal conjugate prior model. The prior is Σ is known and fixed, and it corresponds to the
property |
Note
The multivariate normal hyperparameters μ and V correspond to the
MuandVproperties ofPriorMdl, respectively. The Minnesota prior regularization options [1] enable you to specify μ and V for coefficient shrinkage and tightness completely and easily. You can also display or adjust their values directly by using dot notation afterbayesvarmreturnsPriorMdl.The prior model type that you choose depends on your assumptions about the joint distribution of the parameters. Your choice can affect posterior estimates and inferences. For more details, see Implement Bayesian Linear Regression.
Example: ModelType="conjugate"
Data Types: char | string
Response series names for display, specified as a length m
string vector or cell vector of character vectors. The default is ["Y1" "Y2"
... "Y.m"]
Example: SeriesNames=["CPI" "Unemployment"]
Data Types: string | char
Flag for including a model constant c, specified as a value in this table.
| Value | Description |
|---|---|
false | Response equations do not include a model constant. |
true | All response equations contain a model constant. |
Example: IncludeConstant=false
Data Types: logical
Flag for including a linear time trend term δ, specified as a value in this table.
| Value | Description |
|---|---|
false | Response equations do not include a linear time trend term. |
true | All response equations contain a linear time trend term. |
Example: IncludeTrend=true
Data Types: logical
Number of exogenous predictor variables in the model regression component,
specified as a nonnegative integer. bayesvarm includes all
predictor variables symmetrically in each response equation.
Example: NumPredictors=3
Model description, specified as a string scalar or character vector. The default
value describes the parametric form of the model, for example "2-Dimensional
VAR(3) Model".
Example: Description="Model 1"
Data Types: string | char
Minnesota Prior Regularization Options for Nondiffuse Priors
Center of shrinkage for lag 1 self lags, or the prior expectation on the diagonal
elements of Φ1, specified as a
numseries-by-1 numeric vector.
Center( is the prior mean of .j)
Each element can be any real number, but typical values are in the interval [0,1]. This table describes the prior model of an individual response series for the specified value.
| Value | Prior Model |
|---|---|
0 | White noise process |
In the interval (0,1) | Stationary AR(1) |
1 | Random walk |
bayesvarm sets the prior means of the following variables to 0:
Off-diagonal elements of Φ1
All elements of Φq, q > 1
Model constants c
Linear time trend coefficients δ
Exogenous predictor coefficients Β
For more details, see Minnesota Prior.
Example: Center=0.01*ones(3,1)
Data Types: double
Tightness of shrinkage on all self lags of Φ1, specified as a positive numeric scalar.
SelfLag contributes to the prior variances of all self-lag coefficients in the model (property V of the output model PriorMdl).
Tip
Relatively small tightness values indicate strong belief in prior assumptions during estimation (that is, relatively small values tightly lock self lags around their prior mean). Relatively large values place more weight on information in the data during estimation.
For more details, see Minnesota Prior.
Example: SelfLag=0.5
Data Types: double
Tightness on all cross-variable lag coefficients of Φ1,
specified as a positive numeric scalar. For conjugate prior models,
bayesvarm sets 'CrossLag' to the value of
the SelfLag name-value argument.
CrossLag contributes to the prior variances of all cross-variable lag coefficients in the model (property V of the output model PriorMdl).
Tip
Relatively small tightness values indicate strong belief in prior assumptions during estimation (that is, relatively small values tightly lock cross lags around their prior mean). Relatively large values place more weight on information in the data during estimation.
For more details, see Minnesota Prior.
Example: CrossLag=0.05
Data Types: double
Speed of the prior variance decay with increasing lag, specified as a positive numeric scalar.
Decay contributes to the prior variance of all lag coefficient matrices greater than lag 1 (property V of the output model PriorMdl).
Tip
Relatively large values cause lag coefficient variances to decay more quickly, which tightly locks higher-order lag coefficients to their prior means.
Example: Decay=2
Data Types: double
Response variable variances for the cross-variable lag coefficient tightness
CrossLag, specified as a numseries-by-1
positive numeric vector. Elements correspond to the response variables. For conjugate
prior models, bayesvarm ignores
Scale.
Scale contributes to the prior variances of all cross-variable lag coefficients in the model (property V of the output model PriorMdl), but does not directly contribute to the innovations covariance matrix stored in the property Sigma.
Tip
Specify 'Scale' when response variable scales are unbalanced.
Example: Scale=[2 1]
Data Types: double
Prior variance of exogenous coefficients, specified as a positive numeric scalar.
VarianceX sets the prior variances of all exogenous variables,
including the model constant c, linear time trend term
δ, and exogenous predictor coefficients Β.
VarianceX contributes to the value of the prior coefficient variance (property V of the output model PriorMdl).
Tip
Relatively small tightness values indicate strong belief in prior assumptions during estimation (that is, relatively small values tightly lock coefficients of exogenous variables to their prior means). Relatively large values place more weight on information in the data during estimation.
Example: VarianceX=100
Data Types: double
Innovations Covariance Hyperparameter Options
Fixed innovations covariance matrix for the normal prior model, specified as a
numseries-by-numseries positive definite
numeric matrix.
If you specify ModelType="normal", you must specify
Sigma. For other prior models, Σ is a random variable, so
Sigma does not apply.
Example: Sigma=eye(2)
Data Types: double
Inverse Wishart scale matrix, specified as a
numseries-by-numseries positive definite
numeric matrix.
Example: Omega=eye(numseries)
Data Types: double
Inverse Wishart degrees of freedom, specified as a positive numeric scalar.
For a proper distribution, specify a value that is greater than numseries – 1. For a distribution with a finite mean, specify a value that is greater than numseries + 1.
Example: DoF=8
Data Types: double
Output Arguments
Bayesian VAR model storing prior model assumptions, returned as one of the model objects listed in this table.
Value of ModelType | Returned Bayesian VAR Model Object |
|---|---|
'conjugate' | conjugatebvarm |
'semiconjugate' | semiconjugatebvarm |
'diffuse' | diffusebvarm |
'normal' | normalbvarm |
PriorMdl specifies the joint prior distribution and characteristics of the VAR model only. The model object is a template intended for further use. To incorporate data into the model for posterior distribution analysis, pass the model object and data to the appropriate object function, for example, estimate or simulate.
More About
A Bayesian VAR model treats all coefficients and the innovations covariance matrix as random variables in the m-dimensional, stationary VARX(p) model. The model has one of the three forms described in this table.
| Model | Equation |
|---|---|
| Reduced-form VAR(p) in difference-equation notation |
|
| Multivariate regression |
|
| Matrix regression |
|
For each time t = 1,...,T:
yt is the m-dimensional observed response vector, where m =
numseries.Φ1,…,Φp are the m-by-m AR coefficient matrices of lags 1 through p, where p =
numlags.c is the m-by-1 vector of model constants if
IncludeConstantistrue.δ is the m-by-1 vector of linear time trend coefficients if
IncludeTrendistrue.Β is the m-by-r matrix of regression coefficients of the r-by-1 vector of observed exogenous predictors xt, where r =
NumPredictors. All predictor variables appear in each equation.which is a 1-by-(mp + r + 2) vector, and Zt is the m-by-m(mp + r + 2) block diagonal matrix
where 0z is a 1-by-(mp + r + 2) vector of zeros.
, which is an (mp + r + 2)-by-m random matrix of the coefficients, and the m(mp + r + 2)-by-1 vector λ = vec(Λ).
εt is an m-by-1 vector of random, serially uncorrelated, multivariate normal innovations with the zero vector for the mean and the m-by-m matrix Σ for the covariance. This assumption implies that the data likelihood is
where f is the m-dimensional multivariate normal density with mean ztΛ and covariance Σ, evaluated at yt.
Before considering the data, you impose a joint prior
distribution assumption on (Λ,Σ) (see the ModelType name-value
argument). bayesvarm enables you to adjust hyperparameters by using
the Minnesota prior
assumptions and parameter structure [1]; the structure regularizes
the coefficients. In a Bayesian analysis, the distribution of the parameters is updated with
information about the parameters obtained from the data likelihood. The result is the
joint posterior distribution of (Λ,Σ).
The Minnesota prior, introduced in [1], is a hyperparameter structure for the joint prior distribution of (Λ,Σ) used to obtain a parsimonious model by regularizing the endogenous coefficient matrices of a Bayesian VAR(p) model. Minnesota regularization considers a tuning parameter for the center of shrinkage and several tuning parameters for the tightness of shrinkage.
The center of shrinkage is specified by the prior mean of the coefficients (see the
Center name-value argument). The Minnesota regularization method
sets the prior mean of all coefficients to 0 except the lag 1 self lags (diagonal elements
of the AR coefficient matrix Φ1). The prior mean of each lag 1 self
lag is a real number, typically in the interval [0,1], where (a priori)
response series j is one of the following:
White noise process if prior mean j is 0
AR(1) model if prior mean j is in the interval (0,1)
Random walk if prior mean j is 1
For example, suppose numseries is 2,
numlags is 2, NumPredictors
is 1, and all other model options have default values. If you specify
Center=0.01*ones(2,1), the vectorized prior mean of Λ|Σ is
where
ϕq,j:
is row j of Φq. MATLAB® stores μ in the Mu property of
PriorMdl. You can adjust Mu by using dot
notation.
The tightness of shrinkage is specified by the prior variance of the coefficients ϕr,jk. For all prior models except conjugate,
where:
v0 is the tightness on the prior means of all self lags of Φ1 (
SelfLag).d is the speed of tightness decay (
Decay).is the tightness on the prior means of all cross-variable lag coefficients of Φ1 (
CrossLag).is the prior response variance (element j of
Scale).
For conjugate prior models,
Tips
Because MATLAB does not adjust input data for variable scales, a best practice is to adjust all series to have a similar magnitude. Consequently, the scales of the coefficients are similar.
By default,
bayesvarmcreates Bayesian VAR models by using the Minnesota prior assumptions and parameter structure [1]. After you create a model, you can inspect the effect of coefficient shrinkage by callingsummarize(PriorMdl). You can change the prior mean and variance by settingPriorMdl.MuandPriorMdl.V, respectively.
Alternative Functionality
To create a non-Bayesian VAR model, see varm.
References
[1] Litterman, Robert B. "Forecasting with Bayesian Vector Autoregressions: Five Years of Experience." Journal of Business and Economic Statistics 4, no. 1 (January 1986): 25–38. https://doi.org/10.2307/1391384.
Version History
Introduced in R2020a
See Also
Objects
Functions
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)