TreeBagger
Ensemble of bagged decision trees
Description
A TreeBagger object is an ensemble of bagged decision trees for
either classification or regression. Individual decision trees tend to overfit.
Bagging, which stands for bootstrap aggregation, is an ensemble method that
reduces the effects of overfitting and improves generalization.
Creation
The TreeBagger function grows every tree in the
TreeBagger ensemble model using bootstrap samples of the input data.
Observations not included in a sample are considered "out-of-bag" for that tree. The function
selects a random subset of predictors for each decision split by using the random forest
algorithm [1].
Syntax
Description
Tip
By default, the TreeBagger function grows classification decision
trees. To grow regression decision trees, specify the name-value argument
Method as "regression".
Mdl = TreeBagger(NumTrees,Tbl,ResponseVarName)Mdl) of NumTrees bagged
classification trees, trained by the predictors in the table Tbl and the
class labels in the variable Tbl.ResponseVarName.
Mdl = TreeBagger(NumTrees,Tbl,formula)Mdl trained by the predictors in the table
Tbl. The input formula is an explanatory model of
the response and a subset of predictor variables in Tbl used to fit
Mdl. Specify formula using Wilkinson Notation.
Mdl = TreeBagger(___,Name=Value)Mdl with additional options specified by one or more name-value
arguments, using any of the previous input argument combinations. For example, you can specify
the algorithm used to find the best split on a categorical predictor by using the name-value
argument PredictorSelection.
Input Arguments
Number of decision trees in the bagged ensemble, specified as a positive integer.
Data Types: single | double
Sample data used to train the model, specified as a table. Each row of
Tbl corresponds to one observation, and each column corresponds to one
predictor variable. Optionally, Tbl can contain one additional column for
the response variable. Multicolumn variables and cell arrays other than cell arrays of
character vectors are not allowed.
If
Tblcontains the response variable, and you want to use all remaining variables inTblas predictors, then specify the response variable by usingResponseVarName.If
Tblcontains the response variable, and you want to use only a subset of the remaining variables inTblas predictors, then specify a formula by usingformula.If
Tbldoes not contain the response variable, then specify a response variable by usingY. The length of the response variable and the number of rows inTblmust be equal.
Response variable name, specified as the name of a variable in
Tbl.
You must specify ResponseVarName as a character vector or string
scalar. For example, if the response variable Y is stored as
Tbl.Y, then specify it as "Y". Otherwise, the software
treats all columns of Tbl, including Y, as predictors
when training the model.
The response variable must be a categorical, character, or string array; a logical or
numeric vector; or a cell array of character vectors. If Y is a character
array, then each element of the response variable must correspond to one row of the
array.
A good practice is to specify the order of the classes by using the
ClassNames name-value argument.
Data Types: char | string
Explanatory model of the response variable and a subset of the predictor variables,
specified as a character vector or string scalar in the form "Y~x1+x2+x3".
In this form, Y represents the response variable, and
x1, x2, and x3 represent the
predictor variables.
To specify a subset of variables in Tbl as predictors for training
the model, use a formula. If you specify a formula, then the software does not use any
variables in Tbl that do not appear in
formula.
The variable names in the formula must be both variable names in Tbl
(Tbl.Properties.VariableNames) and valid MATLAB® identifiers. You can verify the variable names in Tbl by
using the isvarname function. If the variable names
are not valid, then you can convert them by using the matlab.lang.makeValidName function.
Data Types: char | string
Class labels or response variable to which the ensemble of bagged decision trees is trained, specified as a categorical, character, or string array; a logical or numeric vector; or a cell array of character vectors.
If you specify
Methodas"classification", the following apply for the class labelsY:Each element of
Ydefines the class membership of the corresponding row ofX.If
Yis a character array, then each row must correspond to one class label.The
TreeBaggerfunction converts the class labels to a cell array of character vectors.
If you specify
Methodas"regression", the response variableYis an n-by-1 numeric vector, where n is the number of observations. Each entry inYis the response for the corresponding row ofX.
The length of Y and the number of rows of X must
be equal.
Data Types: categorical | char | string | logical | single | double | cell
Predictor data, specified as a numeric matrix.
Each row of X corresponds to one observation (also known as an
instance or example), and each column corresponds to one variable (also known as a
feature).
The length of Y and the number of rows of X must
be equal.
Data Types: double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: TreeBagger(100,X,Y,Method="regression",Surrogate="on",OOBPredictorImportance="on")
creates a bagged ensemble of 100 regression trees, and specifies to use surrogate splits and to
store the out-of-bag information for predictor importance estimation.
Number of observations in each chunk of data, specified as a positive integer. This
option applies only when you use TreeBagger on tall arrays. For more
information, see Extended Capabilities.
Example: ChunkSize=10000
Data Types: single | double
Misclassification cost, specified as a square matrix or structure.
If you specify the square matrix
Costand the true class of an observation isi, thenCost(i,j)is the cost of classifying a point into classj. That is, rows correspond to the true classes and columns correspond to the predicted classes. To specify the class order for the corresponding rows and columns ofCost, use theClassNamesname-value argument.If you specify the structure
S, then it must have two fields:S.ClassNames, which contains the class names as a variable of the same data type asYS.ClassificationCosts, which contains the cost matrix with rows and columns ordered as inS.ClassNames
The default value is Cost(i,j)=1 if i~=j, and
Cost(i,j)=0 if i=j.
For more information on the effect of a highly skewed Cost, see
Algorithms.
Example: Cost=[0,1;2,0]
Data Types: single | double | struct
Categorical predictors list, specified as one of the values in this table.
| Value | Description |
|---|---|
| Vector of positive integers |
Each entry in the vector is an index value indicating that the corresponding predictor is
categorical. The index values are between 1 and If |
| Logical vector |
A |
| Character matrix | Each row of the matrix is the name of a predictor variable. The names must match the entries in PredictorNames. Pad the names with extra blanks so each row of the character matrix has the same length. |
| String array or cell array of character vectors | Each element in the array is the name of a predictor variable. The names must match the entries in PredictorNames. |
"all" | All predictors are categorical. |
By default, if the
predictor data is in a table (Tbl), TreeBagger
assumes that a variable is categorical if it is a logical vector, categorical vector, character
array, string array, or cell array of character vectors. If the predictor data is a matrix
(X), TreeBagger assumes that all predictors are
continuous. To identify any other predictors as categorical predictors, specify them by using
the CategoricalPredictors name-value argument.
For the identified categorical predictors, TreeBagger creates
dummy variables using two different schemes, depending on whether a categorical variable is
unordered or ordered. For an unordered categorical variable,
TreeBagger creates one dummy variable for each level of the
categorical variable. For an ordered categorical variable, TreeBagger
creates one less dummy variable than the number of categories. For details, see Automatic Creation of Dummy Variables.
Example: CategoricalPredictors="all"
Data Types: single | double | logical | char | string | cell
Type of decision tree, specified as "classification" or
"regression". For regression trees, Y must be
numeric.
Example: Method="regression"
Minimum number of leaf node observations, specified as a positive integer. Each leaf
has at least MinLeafSize observations per tree leaf. By default,
MinLeafSize is 1 for classification trees and
5 for regression trees.
Example: MinLeafSize=4
Data Types: single | double
Number of predictor variables (randomly selected) for each decision split, specified as
a positive integer or "all". By default,
NumPredictorsToSample is the square root of the number of variables
for classification trees, and one third of the number of variables for regression trees. If
the default number is not an integer, the software rounds the number to the nearest integer
in the direction of positive infinity. If you set NumPredictorsToSample
to any value except "all", the software uses Breiman's random forest
algorithm [1].
Example: NumPredictorsToSample=5
Data Types: single | double | char | string
Number of grown trees (training cycles) after which the software displays a message about the training progress in the command window, specified as a nonnegative integer. By default, the software displays no diagnostic messages.
Example: NumPrint=10
Data Types: single | double
Fraction of input data to sample with replacement from the input data for growing each new tree, specified as a positive scalar in the range (0,1].
Example: InBagFraction=0.5
Data Types: single | double
Indicator to store out-of-bag information in the ensemble, specified as
"on" or "off". Specify
OOBPrediction as "on" to store information on which
observations are out-of-bag for each tree. TreeBagger can use this
information to compute the predicted class probabilities for each tree in the
ensemble.
Example: OOBPrediction="off"
Indicator to store out-of-bag estimates of feature importance in the ensemble,
specified as "on" or "off". If you specify
OOBPredictorImportance as "on", the
TreeBagger function sets OOBPrediction to
"on". If you want to analyze predictor importance, specify
PredictorSelection as "curvature" or
"interaction-curvature".
Example: OOBPredictorImportance="on"
Options for computing in parallel and setting random streams, specified as a
structure. Create the Options structure using statset. This table lists the option fields and their
values.
| Field Name | Value | Default |
|---|---|---|
UseParallel | Set this value to true to run computations in
parallel. | false |
UseSubstreams | Set this value to To compute
reproducibly, set | false |
Streams | Specify this value as a RandStream object or
cell array of such objects. Use a single object except when the
UseParallel value is true
and the UseSubstreams value is
false. In that case, use a cell array that
has the same size as the parallel pool. | If you do not specify Streams, then
TreeBagger uses the default stream or
streams. |
Note
You need Parallel Computing Toolbox™ to run computations in parallel.
Example: Options=statset(UseParallel=true,UseSubstreams=true,Streams=RandStream("mlfg6331_64"))
Data Types: struct
Predictor variable names, specified as a string array of unique names or cell array of
unique character vectors. The functionality of PredictorNames depends
on how you supply the training data.
If you supply
XandY, then you can usePredictorNamesto assign names to the predictor variables inX.The order of the names in
PredictorNamesmust correspond to the column order ofX. That is,PredictorNames{1}is the name ofX(:,1),PredictorNames{2}is the name ofX(:,2), and so on. Also,size(X,2)andnumel(PredictorNames)must be equal.By default,
PredictorNamesis{'x1','x2',...}.
If you supply
Tbl, then you can usePredictorNamesto choose which predictor variables to use in training. That is,TreeBaggeruses only the predictor variables inPredictorNamesand the response variable during training.PredictorNamesmust be a subset ofTbl.Properties.VariableNamesand cannot include the name of the response variable.By default,
PredictorNamescontains the names of all predictor variables.A good practice is to specify the predictors for training using either
PredictorNamesorformula, but not both.
Example: PredictorNames=["SepalLength","SepalWidth","PetalLength","PetalWidth"]
Data Types: string | cell
Indicator for sampling with replacement, specified as "on" or
"off". Specify SampleWithReplacement as
"on" to sample with replacement, or as "off" to
sample without replacement. If you set SampleWithReplacement to
"off", you must set the name-value argument
InBagFraction to a value less than 1.
Example: SampleWithReplacement="on"
Prior probability for each class for two-class learning, specified as a value in this table.
| Value | Description |
|---|---|
"empirical" | The class prior probabilities are the class relative frequencies in
Y. |
"uniform" | All class prior probabilities are equal to 1/K, where K is the number of classes. |
| numeric vector | Each element in the vector is a class prior probability. Order the elements
according to Mdl.ClassNames, or specify the order using the
ClassNames name-value argument. The software normalizes the
elements to sum to 1. |
| structure | A structure
|
If you specify a cost matrix, the Prior property of the
TreeBagger model stores the prior probabilities adjusted for the
misclassification cost. For more details, see Algorithms.
This argument is valid only for two-class learning.
Example: Prior=struct(ClassNames=["setosa" "versicolor"
"virginica"],ClassProbs=1:3)
Data Types: char | string | single | double | struct
Note
In addition to its name-value arguments, the TreeBagger function
accepts the name-value arguments of fitctree and fitrtree listed in Additional Name-Value Arguments of TreeBagger Function.
Output Arguments
Ensemble of bagged decision trees, returned as a TreeBagger
object.
Properties
Bagging Properties
This property is read-only.
Indicator to compute out-of-bag predictions for training observations, specified as a
numeric or logical 1 (true) or 0 (false). If this
property is true:
The
TreeBaggerobject has the propertiesOOBIndicesandOOBInstanceWeight.You can use the object functions
oobError,oobMargin, andoobMeanMargin.
This property is read-only.
Indicator to compute the out-of-bag variable importance, specified as a numeric or
logical 1 (true) or 0 (false). If this property is
true:
The
TreeBaggerobject has the propertiesOOBPermutedPredictorDeltaError,OOBPermutedPredictorDeltaMeanMargin, andOOBPermutedPredictorCountRaiseMargin.The property
ComputeOOBPredictionis alsotrue.
This property is read-only.
Fraction of observations that are randomly selected with replacement (in-bag
observations) for each bootstrap replica, specified as a numeric scalar. The size of each
replica is Nobs×InBagFraction, where
Nobs is the number of observations in the training data.
Data Types: single | double
This property is read-only.
Out-of-bag indices, specified as a logical array. This property is a
Nobs-by-NumTrees array, where
Nobs is the number of observations in the training data, and
NumTrees is the number of trees in the ensemble. If the
OOBIndices(i,j)true, the observation i is out-of-bag for
the tree j (that is, the TreeBagger function did not
select the observation i for the training data used to grow the tree
j).
This property is read-only.
Number of out-of-bag trees for each observation, specified as a numeric vector. This
property is a Nobs-by-1 vector, where Nobs is the
number of observations in the training data. The
OOBInstanceWeight(i)
Data Types: single | double
This property is read-only.
Predictor variable (feature) importance for raising the margin, specified as a numeric vector. This property is a 1-by-Nvars vector, where Nvars is the number of variables in the training data. For each variable, the measure is the difference between the number of raised margins and the number of lowered margins if the values of that variable are permuted across the out-of-bag observations. This measure is computed for every tree, then averaged over the entire ensemble and divided by the standard deviation over the entire ensemble.
This property is empty ([]) for regression trees.
Data Types: single | double
This property is read-only.
Predictor variable (feature) importance for prediction error, specified as a numeric vector. This property is a 1-by-Nvars vector, where Nvars is the number of variables (columns) in the training data. For each variable, the measure is the increase in prediction error if the values of that variable are permuted across the out-of-bag observations. This measure is computed for every tree, then averaged over the entire ensemble and divided by the standard deviation over the entire ensemble.
Data Types: single | double
This property is read-only.
Predictor variable (feature) importance for the classification margin, specified as numeric vector. This property is a 1-by-Nvars vector, where Nvars is the number of variables (columns) in the training data. For each variable, the measure is the decrease in the classification margin if the values of that variable are permuted across the out-of-bag observations. This measure is computed for every tree, then averaged over the entire ensemble and divided by the standard deviation over the entire ensemble.
This property is empty ([]) for regression trees.
Data Types: single | double
Tree Properties
This property is read-only.
Split criterion contributions for each predictor, specified as a numeric vector. This property is a 1-by-Nvars vector, where Nvars is the number of changes in the split criterion. The software sums the changes in the split criterion over splits on each variable, then averages the sums across the entire ensemble of grown trees.
Data Types: single | double
This property is read-only.
Indicator to merge leaves, specified as a numeric or logical 1 (true)
or 0 (false). This property is true if the software
merges the decision tree leaves with the same parent, for splits that do not decrease the
total risk. Otherwise, this property is false.
This property is read-only.
Minimum number of leaf node observations, specified as a positive integer. Each leaf has
at least MinLeafSize observations. By default,
MinLeafSize is 1 for classification trees and 5 for regression trees.
For decision tree training, fitctree and fitrtree
set the name-value argument MinParentSize to
2*MinLeafSize.
Data Types: single | double
This property is read-only.
Number of decision trees in the bagged ensemble, specified as a positive integer.
Data Types: single | double
This property is read-only.
Indicator to estimate the optimal sequence of pruned subtrees, specified as a numeric
or logical 1 (true) or 0 (false). The
Prune property is true if the decision trees are
pruned, and false if they are not. Pruning decision trees is not
recommended for ensembles.
This property is read-only.
Indicator to sample each decision tree with replacement, specified as a numeric or
logical 1 (true) or 0 (false). This property is
true if the TreeBagger function samples each
decision tree with replacement, and false otherwise.
This property is read-only.
Predictive measures of variable association, specified as a numeric matrix. This property is an Nvars-by-Nvars matrix, where Nvars is the number of predictor variables. The property contains the predictive measures of variable association, averaged across the entire ensemble of grown trees.
If you grow the ensemble with the
Surrogatename-value argument set to"on", this matrix, for each tree, is filled with the predictive measures of association averaged over the surrogate splits.If you grow the ensemble with the
Surrogatename-value argument set to"off", theSurrogateAssociationproperty is an identity matrix. By default,Surrogateis set to"off".
Data Types: single | double
This property is read-only.
Name-value arguments specified for the TreeBagger function,
specified as a cell array. The TreeBagger function uses these name-value
arguments when it grows new trees for the bagged ensemble.
This property is read-only.
Decision trees in the bagged ensemble, specified as a NumTrees-by-1 cell
array. Each tree is a CompactClassificationTree or
CompactRegressionTree object.
Predictor Properties
This property is read-only.
Number of decision splits for each predictor, specified as a numeric vector. This property is
a 1-by-Nvars vector, where
Nvars is the number of predictor
variables. Each element of
NumPredictorSplit represents
the number of splits on the predictor summed over all
trees.
Data Types: single | double
This property is read-only.
Number of predictor variables to select at random for each decision split, specified as a positive integer. By default, this property is the square root of the total number of variables for classification trees, and one third of the total number of variables for regression trees.
Data Types: single | double
This property is read-only.
Outlier measure for each observation, specified as a numeric vector. This property is a Nobs-by-1 vector, where Nobs is the number of observations in the training data.
Data Types: single | double
This property is read-only.
Predictor names, specified as a cell array of character vectors. The order of the elements in PredictorNames corresponds to the order in which the predictor names appear in the training data X.
This property is read-only.
Predictors used to train the bagged ensemble, specified as a numeric array. This property is a Nobs-by-Nvars array, where Nobs is the number of observations (rows) and Nvars is the number of variables (columns) in the training data.
Data Types: single | double
Response Properties
Default prediction value returned by predict or
oobPredict, specified as "",
"MostPopular", or a numeric scalar. This property controls the predicted
value returned by the predict or oobPredict object
function when no prediction is possible (for example, when oobPredict
predicts a response for an observation that is in-bag for all trees in the ensemble).
For classification trees, you can set
DefaultYfitto either""or"MostPopular". If you specify"MostPopular"(default for classification), the property value is the name of the most probable class in the training data. If you specify"", the in-bag observations are excluded from computation of the out-of-bag error and margin.For regression trees, you can set
DefaultYfitto any numeric scalar. The default value for regression is the mean of the response for the training data. If you setDefaultYfittoNaN, the in-bag observations are excluded from computation of the out-of-bag error and margin.
Example: Mdl.DefaultYfit="MostPopular"
Data Types: single | double | char | string
This property is read-only.
Class labels or response data, specified as a cell array of character vectors or a numeric vector.
If you set the
Methodname-value argument to"classification", this property represents class labels. Each row ofYrepresents the observed classification of the corresponding row ofX.If you set the
Methodname-value argument to"regression", this property represents response data and is a numeric vector.
Data Types: single | double | cell
Training Properties
This property is read-only.
Type of ensemble, specified as "classification" for classification
ensembles or "regression" for regression ensembles.
This property is read-only.
Proximity between training data observations, specified as a numeric array. This property is a Nobs-by-Nobs array, where Nobs is the number of observations in the training data. The array contains measures of the proximity between observations. For any two observations, their proximity is defined as the fraction of trees for which these observations land on the same leaf. The array is symmetric, with ones on the diagonal and off-diagonal elements ranging from 0 to 1.
Data Types: single | double
This property is read-only.
Observation weights, specified as a vector of nonnegative values. This property has the
same number of rows as Y. Each entry in W specifies
the relative importance of the corresponding observation in Y. The
TreeBagger function uses the observation weights to grow each decision
tree in the ensemble.
Data Types: single | double
Classification Properties
This property is read-only.
Unique class names used in the training model, specified as a cell array of character vectors.
This property is empty ([]) for regression trees.
This property is read-only.
Misclassification cost, specified as a numeric square matrix. The element
Cost(i,j) is the cost of classifying a point into class
j if its true class is i. The rows correspond to the
true class and the columns correspond to the predicted class. The order of the rows and
columns of Cost corresponds to the order of the classes in
ClassNames.
This property is empty ([]) for regression trees.
Data Types: single | double
This property is read-only.
Prior probabilities, specified as a numeric vector. The order of the elements in
Prior corresponds to the order of the elements in
Mdl.ClassNames.
If you specify a cost matrix by using the Cost name-value argument
of the TreeBagger function, the Prior property of the
TreeBagger model object stores the prior probabilities (specified by the
Prior name-value argument) adjusted for the misclassification cost. For
more details, see Algorithms.
This property is empty ([]) for regression trees.
Data Types: single | double
Object Functions
compact | Compact ensemble of decision trees |
partialDependence | Compute partial dependence |
plotPartialDependence | Create partial dependence plot (PDP) and individual conditional expectation (ICE) plots |
error | Error (misclassification probability or MSE) |
meanMargin | Mean classification margin |
margin | Classification margin |
oobError | Out-of-bag error |
oobMeanMargin | Out-of-bag mean margins |
oobMargin | Out-of-bag margins |
oobQuantileError | Out-of-bag quantile loss of bag of regression trees |
quantileError | Quantile loss using bag of regression trees |
oobPredict | Ensemble predictions for out-of-bag observations |
oobQuantilePredict | Quantile predictions for out-of-bag observations from bag of regression trees |
predict | Predict responses using ensemble of bagged decision trees |
quantilePredict | Predict response quantile using bag of regression trees |
Examples
Create an ensemble of bagged classification trees for Fisher's iris data set. Then, view the first grown tree, plot the out-of-bag classification error, and predict labels for out-of-bag observations.
Load the fisheriris data set. Create X as a numeric matrix that contains four measurements for 150 irises. Create Y as a cell array of character vectors that contains the corresponding iris species.
load fisheriris
X = meas;
Y = species;Set the random number generator to default for reproducibility.
rng("default")Train an ensemble of bagged classification trees using the entire data set. Specify 50 weak learners. Store the out-of-bag observations for each tree. By default, TreeBagger grows deep trees.
Mdl = TreeBagger(50,X,Y,... Method="classification",... OOBPrediction="on")
Mdl =
TreeBagger
Ensemble with 50 bagged decision trees:
Training X: [150x4]
Training Y: [150x1]
Method: classification
NumPredictors: 4
NumPredictorsToSample: 2
MinLeafSize: 1
InBagFraction: 1
SampleWithReplacement: 1
ComputeOOBPrediction: 1
ComputeOOBPredictorImportance: 0
Proximity: []
ClassNames: 'setosa' 'versicolor' 'virginica'
Properties, Methods
Mdl is a TreeBagger ensemble for classification trees.
The Mdl.Trees property is a 50-by-1 cell vector that contains the trained classification trees for the ensemble. Each tree is a CompactClassificationTree object. View the graphical display of the first trained classification tree.
view(Mdl.Trees{1},Mode="graph")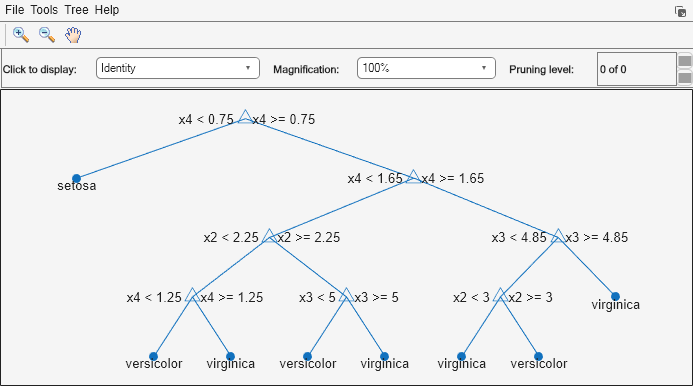
Plot the out-of-bag classification error over the number of grown classification trees.
plot(oobError(Mdl)) xlabel("Number of Grown Trees") ylabel("Out-of-Bag Classification Error")
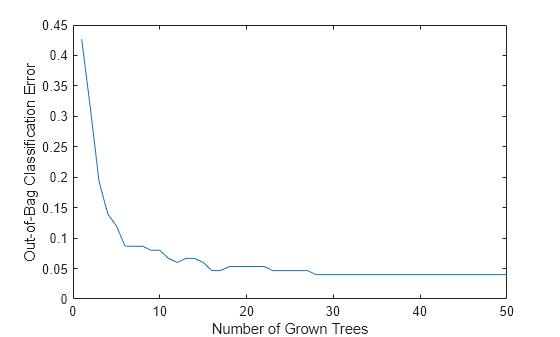
The out-of-bag error decreases as the number of grown trees increases.
Predict labels for out-of-bag observations. Display the results for a random set of 10 observations.
oobLabels = oobPredict(Mdl); ind = randsample(length(oobLabels),10); table(Y(ind),oobLabels(ind),... VariableNames=["TrueLabel" "PredictedLabel"])
ans=10×2 table
TrueLabel PredictedLabel
______________ ______________
{'setosa' } {'setosa' }
{'virginica' } {'virginica' }
{'setosa' } {'setosa' }
{'virginica' } {'virginica' }
{'setosa' } {'setosa' }
{'virginica' } {'virginica' }
{'setosa' } {'setosa' }
{'versicolor'} {'versicolor'}
{'versicolor'} {'virginica' }
{'virginica' } {'virginica' }
Create an ensemble of bagged regression trees for the carsmall data set. Then, predict conditional mean responses and conditional quartiles.
Load the carsmall data set. Create X as a numeric vector that contains the car engine displacement values. Create Y as a numeric vector that contains the corresponding miles per gallon.
load carsmall
X = Displacement;
Y = MPG;Set the random number generator to default for reproducibility.
rng("default")Train an ensemble of bagged regression trees using the entire data set. Specify 100 weak learners.
Mdl = TreeBagger(100,X,Y,... Method="regression")
Mdl =
TreeBagger
Ensemble with 100 bagged decision trees:
Training X: [94x1]
Training Y: [94x1]
Method: regression
NumPredictors: 1
NumPredictorsToSample: 1
MinLeafSize: 5
InBagFraction: 1
SampleWithReplacement: 1
ComputeOOBPrediction: 0
ComputeOOBPredictorImportance: 0
Proximity: []
Properties, Methods
Mdl is a TreeBagger ensemble for regression trees.
For 10 equally spaced engine displacements between the minimum and maximum in-sample displacement, predict conditional mean responses (YMean) and conditional quartiles (YQuartiles).
predX = linspace(min(X),max(X),10)';
YMean = predict(Mdl,predX);
YQuartiles = quantilePredict(Mdl,predX,...
Quantile=[0.25,0.5,0.75]);Plot the observations, estimated mean responses, and estimated quartiles.
hold on plot(X,Y,"o"); plot(predX,YMean) plot(predX,YQuartiles) hold off ylabel("Fuel Economy") xlabel("Engine Displacement") legend("Data","Mean Response",... "First Quartile","Median",..., "Third Quartile")
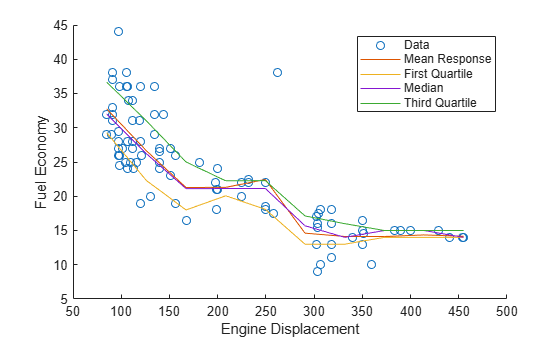
Create two ensembles of bagged regression trees, one using the standard CART algorithm for splitting predictors, and the other using the curvature test for splitting predictors. Then, compare the predictor importance estimates for the two ensembles.
Load the carsmall data set and convert the variables Cylinders, Mfg, and Model_Year to categorical variables. Then, display the number of categories represented in the categorical variables.
load carsmall
Cylinders = categorical(Cylinders);
Mfg = categorical(cellstr(Mfg));
Model_Year = categorical(Model_Year);
numel(categories(Cylinders))ans = 3
numel(categories(Mfg))
ans = 28
numel(categories(Model_Year))
ans = 3
Create a table that contains eight car metrics.
Tbl = table(Acceleration,Cylinders,Displacement,...
Horsepower,Mfg,Model_Year,Weight,MPG);Set the random number generator to default for reproducibility.
rng("default")Train an ensemble of 200 bagged regression trees using the entire data set. Because the data has missing values, specify to use surrogate splits. Store the out-of-bag information for predictor importance estimation.
By default, TreeBagger uses the standard CART, an algorithm for splitting predictors. Because the variables Cylinders and Model_Year each contain only three categories, the standard CART prefers splitting a continuous predictor over these two variables.
MdlCART = TreeBagger(200,Tbl,"MPG",... Method="regression",Surrogate="on",... OOBPredictorImportance="on");
TreeBagger stores predictor importance estimates in the property OOBPermutedPredictorDeltaError.
impCART = MdlCART.OOBPermutedPredictorDeltaError;
Train a random forest of 200 regression trees using the entire data set. To grow unbiased trees, specify to use the curvature test for splitting predictors.
MdlUnbiased = TreeBagger(200,Tbl,"MPG",... Method="regression",Surrogate="on",... PredictorSelection="curvature",... OOBPredictorImportance="on"); impUnbiased = MdlUnbiased.OOBPermutedPredictorDeltaError;
Create bar graphs to compare the predictor importance estimates impCART and impUnbiased for the two ensembles.
tiledlayout(1,2,Padding="compact"); nexttile bar(impCART) title("Standard CART") ylabel("Predictor Importance Estimates") xlabel("Predictors") h = gca; h.XTickLabel = MdlCART.PredictorNames; h.XTickLabelRotation = 45; h.TickLabelInterpreter = "none"; nexttile bar(impUnbiased); title("Curvature Test") ylabel("Predictor Importance Estimates") xlabel("Predictors") h = gca; h.XTickLabel = MdlUnbiased.PredictorNames; h.XTickLabelRotation = 45; h.TickLabelInterpreter = "none";
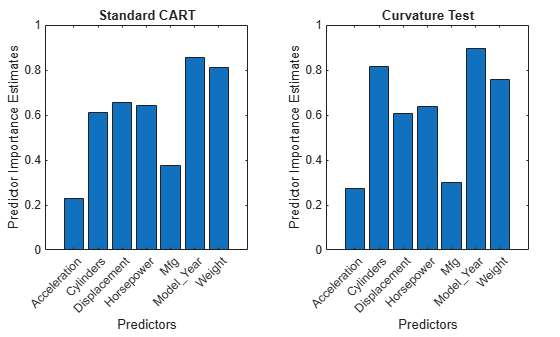
For the CART model, the continuous predictor Weight is the second most important predictor. For the unbiased model, the predictor importance of Weight is smaller in value and ranking.
Train an ensemble of bagged classification trees for observations in a tall array, and find the misclassification probability of each tree in the model for weighted observations. This example uses the data set airlinesmall.csv, a large data set that contains a tabular file of airline flight data.
When you perform calculations on tall arrays, MATLAB® uses either a parallel pool (default if you have Parallel Computing Toolbox™) or the local MATLAB session. To run the example using the local MATLAB session when you have Parallel Computing Toolbox, change the global execution environment by using the mapreducer function.
mapreducer(0)
Create a datastore that references the location of the folder containing the data set. Select a subset of the variables to work with, and treat "NA" values as missing data so that the datastore function replaces them with NaN values. Create the tall table tt to contain the data in the datastore.
ds = datastore("airlinesmall.csv"); ds.SelectedVariableNames = ["Month" "DayofMonth" "DayOfWeek",... "DepTime" "ArrDelay" "Distance" "DepDelay"]; ds.TreatAsMissing = "NA"; tt = tall(ds)
tt =
M×7 tall table
Month DayofMonth DayOfWeek DepTime ArrDelay Distance DepDelay
_____ __________ _________ _______ ________ ________ ________
10 21 3 642 8 308 12
10 26 1 1021 8 296 1
10 23 5 2055 21 480 20
10 23 5 1332 13 296 12
10 22 4 629 4 373 -1
10 28 3 1446 59 308 63
10 8 4 928 3 447 -2
10 10 6 859 11 954 -1
: : : : : : :
: : : : : : :
Determine the flights that are late by 10 minutes or more by defining a logical variable that is true for a late flight. This variable contains the class labels Y. A preview of this variable includes the first few rows.
Y = tt.DepDelay > 10
Y = M×1 tall logical array 1 0 1 1 0 1 0 0 : :
Create a tall array X for the predictor data.
X = tt{:,1:end-1}X =
M×6 tall double matrix
10 21 3 642 8 308
10 26 1 1021 8 296
10 23 5 2055 21 480
10 23 5 1332 13 296
10 22 4 629 4 373
10 28 3 1446 59 308
10 8 4 928 3 447
10 10 6 859 11 954
: : : : : :
: : : : : :
Create a tall array W for the observation weights by arbitrarily assigning double weights to the observations in class 1.
W = Y+1;
Remove the rows in X, Y, and W that contain missing data.
R = rmmissing([X Y W]); X = R(:,1:end-2); Y = R(:,end-1); W = R(:,end);
Train an ensemble of 20 bagged classification trees using the entire data set. Specify a weight vector and uniform prior probabilities. For reproducibility, set the seeds of the random number generators using rng and tallrng. The results can vary depending on the number of workers and the execution environment for the tall arrays. For details, see Control Where Your Code Runs.
rng("default") tallrng("default") tMdl = TreeBagger(20,X,Y,... Weights=W,Prior="uniform")
Evaluating tall expression using the Local MATLAB Session: - Pass 1 of 1: Completed in 0.44 sec Evaluation completed in 0.47 sec Evaluating tall expression using the Local MATLAB Session: - Pass 1 of 1: Completed in 1.5 sec Evaluation completed in 1.6 sec Evaluating tall expression using the Local MATLAB Session: - Pass 1 of 1: Completed in 3.8 sec Evaluation completed in 3.8 sec
tMdl =
CompactTreeBagger
Ensemble with 20 bagged decision trees:
Method: classification
NumPredictors: 6
ClassNames: '0' '1'
Properties, Methods
tMdl is a CompactTreeBagger ensemble with 20 bagged decision trees. For tall data, the TreeBagger function returns a CompactTreeBagger object.
Calculate the misclassification probability of each tree in the model. Attribute a weight contained in the vector W to each observation by using the Weights name-value argument.
terr = error(tMdl,X,Y,Weights=W)
Evaluating tall expression using the Local MATLAB Session: - Pass 1 of 1: Completed in 4.7 sec Evaluation completed in 4.7 sec
terr = 20×1
0.1420
0.1214
0.1115
0.1078
0.1037
0.1027
0.1005
0.0997
0.0981
0.0983
⋮
Find the average misclassification probability for the ensemble of decision trees.
avg_terr = mean(terr)
avg_terr = 0.1022
More About
In addition to its Name-Value Arguments, the
TreeBagger function accepts the following name-value arguments of
fitctree and fitrtree.
Supported fitctree Arguments | Supported fitrtree Arguments |
|---|---|
AlgorithmForCategorical | MaxNumSplits |
ClassNames* | MergeLeaves |
MaxNumCategories | PredictorSelection |
MaxNumSplits | Prune |
MergeLeaves | PruneCriterion |
PredictorSelection | QuadraticErrorTolerance |
Prune | SplitCriterion |
PruneCriterion | Surrogate |
SplitCriterion | Weights |
Surrogate | N/A |
Weights | N/A |
*When you specify the ClassNames name-value argument as a logical
vector, use 0 and 1 values. Do not use false and true
values. For example, you can specify ClassNames as [1 0
1].
Tips
For a
TreeBaggermodelMdl, theTreesproperty contains a cell vector ofMdl.NumTreesCompactClassificationTreeorCompactRegressionTreeobjects. View the graphical display of thetgrown tree by entering:view(Mdl.Trees{t})For regression problems,
TreeBaggersupports mean and quantile regression (that is, quantile regression forest [5]).To predict mean responses or estimate the mean squared error given data, pass a
TreeBaggermodel object and the data topredictorerror, respectively. To perform similar operations for out-of-bag observations, useoobPredictoroobError.To estimate quantiles of the response distribution or the quantile error given data, pass a
TreeBaggermodel object and the data toquantilePredictorquantileError, respectively. To perform similar operations for out-of-bag observations, useoobQuantilePredictoroobQuantileError.
Standard CART tends to select split predictors containing many distinct values, such as continuous variables, over those containing few distinct values, such as categorical variables [4]. Consider specifying the curvature or interaction test if either of the following is true:
The data has predictors with relatively fewer distinct values than other predictors; for example, the predictor data set is heterogeneous.
Your goal is to analyze predictor importance.
TreeBaggerstores predictor importance estimates in theOOBPermutedPredictorDeltaErrorproperty.
For more information on predictor selection, see the name-value argument
PredictorSelectionfor classification trees or the name-value argumentPredictorSelectionfor regression trees.
Algorithms
If you specify the
Cost,Prior, andWeightsname-value arguments, the output model object stores the specified values in theCost,Prior, andWproperties, respectively. TheCostproperty stores the user-specified cost matrix (C) without modification. ThePriorandWproperties store the prior probabilities and observation weights, respectively, after normalization. For model training, the software updates the prior probabilities and observation weights to incorporate the penalties described in the cost matrix. For details, see Misclassification Cost Matrix, Prior Probabilities, and Observation Weights.The
TreeBaggerfunction generates in-bag samples by oversampling classes with large misclassification costs and undersampling classes with small misclassification costs. Consequently, out-of-bag samples have fewer observations from classes with large misclassification costs and more observations from classes with small misclassification costs. If you train a classification ensemble using a small data set and a highly skewed cost matrix, then the number of out-of-bag observations per class might be very low. Therefore, the estimated out-of-bag error might have a large variance and be difficult to interpret. The same phenomenon can occur for classes with large prior probabilities.For details on how the
TreeBaggerfunction selects split predictors, and for information on node-splitting algorithms when the function grows decision trees, see Algorithms for classification trees and Algorithms for regression trees.
Alternative Functionality
Statistics and Machine Learning Toolbox™ offers three objects for bagging and random forest:
ClassificationBaggedEnsembleobject created by thefitcensemblefunction for classificationRegressionBaggedEnsembleobject created by thefitrensemblefunction for regressionTreeBaggerobject created by theTreeBaggerfunction for classification and regression
For details about the differences between TreeBagger and
bagged ensembles (ClassificationBaggedEnsemble and
RegressionBaggedEnsemble), see Comparison of TreeBagger and Bagged Ensembles.
References
[1] Breiman, Leo. "Random Forests." Machine Learning 45 (2001): 5–32. https://doi.org/10.1023/A:1010933404324.
[2] Breiman, Leo, Jerome Friedman, Charles J. Stone, and R. A. Olshen. Classification and Regression Trees. Boca Raton, FL: CRC Press, 1984.
[3] Loh, Wei-Yin. "Regression Trees with Unbiased Variable Selection and Interaction Detection." Statistica Sinica 12, no. 2 (2002): 361–386. https://www.jstor.org/stable/24306967.
[4] Loh, Wei-Yin, and Yu-Shan Shih. "Split Selection for Classification Trees." Statistica Sinica 7, no. 4 (1997): 815–840. https://www.jstor.org/stable/24306157.
[5] Meinshausen, Nicolai. "Quantile Regression Forests." Journal of Machine Learning Research 7, no. 35 (2006): 983–999. https://jmlr.org/papers/v7/meinshausen06a.html.
[6] Genuer, Robin, Jean-Michel Poggi, Christine Tuleau-Malot, and Nathalie Villa-Vialanei. "Random Forests for Big Data." Big Data Research 9 (2017): 28–46. https://doi.org/10.1016/j.bdr.2017.07.003.
Extended Capabilities
This function supports tall arrays with the following limitations.
The
TreeBaggerfunction supports these syntaxes for tallX,Y, andTbl:B = TreeBagger(NumTrees,Tbl,Y)B = TreeBagger(NumTrees,X,Y)B = TreeBagger(___,Name=Value)
For tall arrays, the
TreeBaggerfunction supports classification but not regression.The
TreeBaggerfunction supports these name-value arguments:NumPredictorsToSample— The default value is the square root of the number of variables for classification.MinLeafSize— The default value is1if the number of observations is less than 50,000. If the number of observations is 50,000 or greater, then the default value ismax(1,min(5,floor(0.01*NobsChunk))), whereNobsChunkis the number of observations in a chunk.ChunkSize(only for tall arrays) — The default value is50000.
In addition, the
TreeBaggerfunction supports these name-value arguments offitctree:AlgorithmForCategoricalCategoricalPredictorsCost— The columns of the cost matrixCcannot containInforNaNvalues.MaxNumCategoriesMaxNumSplitsMergeLeavesPredictorNamesPredictorSelectionPriorPrunePruneCriterionSplitCriterionSurrogateWeights
For tall data, the
TreeBaggerfunction returns aCompactTreeBaggerobject that contains most of the same properties as a fullTreeBaggerobject. The main difference is that the compact object is more memory efficient. The compact object does not contain properties that include the data, or that include an array of the same size as the data.The number of trees contained in the returned
CompactTreeBaggerobject can differ from the number of trees specified as input to theTreeBaggerfunction.TreeBaggerdetermines the number of trees to return based on factors that include the size of the input data set and the number of data chunks available to grow trees.Supported
CompactTreeBaggerobject functions are:combineerrormarginmeanMarginpredictsetDefaultYfit
The
error,margin,meanMargin, andpredictobject functions do not support the name-value argumentsTrees,TreeWeights, orUseInstanceForTree. ThemeanMarginfunction also does not support theWeightsname-value argument.The
TreeBaggerfunction creates a random forest by generating trees on disjoint chunks of the data. When more data is available than is required to create the random forest, the function subsamples the data. For a similar example, see Random Forests for Big Data [6].Depending on how the data is stored, some chunks of data might contain observations from only a few classes out of all the classes. In this case, the
TreeBaggerfunction might produce inferior results compared to the case where each chunk of data contains observations from most of the classes.During training of the
TreeBaggeralgorithm, the speed, accuracy, and memory usage depend on a number of factors. These factors include the values forNumTreesand the name-value argumentsChunkSize,MinLeafSize, andMaxNumSplits.For an n-by-p tall array
X,TreeBaggerimplements sampling during training. This sampling depends on these variables:Number of trees
NumTreesChunk size
ChunkSizeNumber of observations n
Number of chunks r (approximately equal to
n/ChunkSize)
Because the value of n is fixed for a given
X, your settings forNumTreesandChunkSizedetermine howTreeBaggersamplesX.If r >
NumTrees, thenTreeBaggersamplesChunkSize * NumTreesobservations fromX, and trains one tree per chunk (with each chunk containingChunkSizenumber of observations). This scenario is the most common when you work with tall arrays.If r ≤
NumTrees, thenTreeBaggertrains approximatelyNumTrees/rtrees in each chunk, using bootstrapping within the chunk.If n ≤
ChunkSize, thenTreeBaggeruses bootstrapping to generate samples (each of size n) on which to train individual trees.
When you specify a value for
NumTrees, consider the following:If you run your code on Apache® Spark™, and your data set is distributed with Hadoop® Distributed File System (HDFS™), start by specifying a value for
NumTreesthat is at least twice the number of partitions in HDFS for your data set. This setting prevents excessive data communication among Apache Spark executors and can improve performance of theTreeBaggeralgorithm.TreeBaggercopies fitted trees into the client memory in the resultingCompactTreeBaggermodel. Therefore, the amount of memory available to the client creates an upper bound on the value you can set forNumTrees. You can tune the values ofMinLeafSizeandMaxNumSplitsfor more efficient speed and memory usage at the expense of some predictive accuracy. After tuning, if the value ofNumTreesis less than twice the number of partitions in HDFS for your data set, then consider repartitioning your data in HDFS to have larger partitions.
After you specify a value for
NumTrees, setChunkSizeto ensure thatTreeBaggeruses most of the data to grow trees. Ideally,ChunkSize * NumTreesshould approximate n, the number of rows in your data. Note that the memory available in the workers for training individual trees can also determine an upper bound forChunkSize.You can adjust the Apache Spark memory properties to avoid out-of-memory errors and support your workflow. See
parallel.cluster.Hadoop(Parallel Computing Toolbox) for more information.
For more information, see Tall Arrays for Out-of-Memory Data.
To run in parallel, specify the Options name-value argument in the call to
this function and set the UseParallel field of the
options structure to true using
statset:
Options=statset(UseParallel=true)
For more information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
Version History
Introduced in R2009aStarting in R2022a, the Cost property stores the user-specified cost
matrix. The software stores normalized prior probabilities (Prior) and
observation weights (W) that do not reflect the penalties described in the
cost matrix.
Note that model training has not changed and, therefore, the decision boundaries between classes have not changed.
For training, the fitting function updates the specified prior probabilities by
incorporating the penalties described in the specified cost matrix, and then normalizes the
prior probabilities and observation weights. This behavior has not changed. In previous
releases, the software stored the default cost matrix in the Cost property
and stored the prior probabilities and observation weights used for training in the
Prior and W properties, respectively. Starting in
R2022a, the software stores the user-specified cost matrix without modification, and stores
normalized prior probabilities and observation weights that do not reflect the cost penalties.
For more details, see Misclassification Cost Matrix, Prior Probabilities, and Observation Weights.
The oobError and oobMeanMargin functions use the
observation weights stored in the W property. Therefore, if you specify a
nondefault cost matrix when you train a classification model, the object functions return a
different value compared to previous releases.
If you want the software to handle the cost matrix, prior
probabilities, and observation weights in the same way as in previous releases, adjust the prior
probabilities and observation weights for the nondefault cost matrix, as described in Adjust Prior Probabilities and Observation Weights for Misclassification Cost Matrix. Then, when you train a
classification model, specify the adjusted prior probabilities and observation weights by using
the Prior and Weights name-value arguments, respectively,
and use the default cost matrix.
See Also
Objects
Functions
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)