resubPredict
Classify training data using trained classifier
Syntax
Description
[
specifies whether to include interaction terms in computations. This syntax applies only
to generalized additive models.label,Score] = resubPredict(Mdl,'IncludeInteractions',includeInteractions)
Examples
Load the fisheriris data set. Create X as a numeric matrix that contains four measurements for 150 irises. Create Y as a cell array of character vectors that contains the corresponding iris species.
load fisheriris X = meas; Y = species; rng('default') % For reproducibility
Train a naive Bayes classifier using the predictors X and class labels Y. A recommended practice is to specify the class names. fitcnb assumes that each predictor is conditionally and normally distributed.
Mdl = fitcnb(X,Y,'ClassNames',{'setosa','versicolor','virginica'})
Mdl =
ClassificationNaiveBayes
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'setosa' 'versicolor' 'virginica'}
ScoreTransform: 'none'
NumObservations: 150
DistributionNames: {'normal' 'normal' 'normal' 'normal'}
DistributionParameters: {3×4 cell}
Properties, Methods
Mdl is a trained ClassificationNaiveBayes classifier.
Predict the training sample labels.
label = resubPredict(Mdl);
Display the results for a random set of 10 observations.
idx = randsample(size(X,1),10); table(Y(idx),label(idx),'VariableNames', ... {'True Label','Predicted Label'})
ans=10×2 table
True Label Predicted Label
______________ _______________
{'virginica' } {'virginica' }
{'setosa' } {'setosa' }
{'virginica' } {'virginica' }
{'versicolor'} {'versicolor'}
{'virginica' } {'virginica' }
{'versicolor'} {'versicolor'}
{'virginica' } {'virginica' }
{'setosa' } {'setosa' }
{'virginica' } {'virginica' }
{'setosa' } {'setosa' }
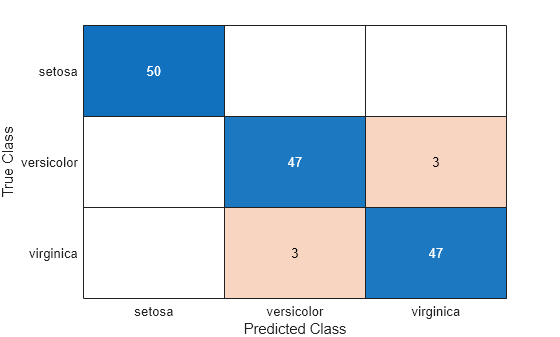
Create a confusion chart from the true labels Y and the predicted labels label.
cm = confusionchart(Y,label);
Load the ionosphere data set. This data set has 34 predictors and 351 binary responses for radar returns, either bad ('b') or good ('g').
load ionosphereTrain a support vector machine (SVM) classifier. Standardize the data and specify that 'g' is the positive class.
SVMModel = fitcsvm(X,Y,'ClassNames',{'b','g'},'Standardize',true);
SVMModel is a ClassificationSVM classifier.
Fit the optimal score-to-posterior-probability transformation function.
rng(1); % For reproducibility
ScoreSVMModel = fitPosterior(SVMModel)ScoreSVMModel =
ClassificationSVM
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'b' 'g'}
ScoreTransform: '@(S)sigmoid(S,-9.482430e-01,-1.217774e-01)'
NumObservations: 351
Alpha: [90×1 double]
Bias: -0.1342
KernelParameters: [1×1 struct]
Mu: [0.8917 0 0.6413 0.0444 0.6011 0.1159 0.5501 0.1194 0.5118 0.1813 0.4762 0.1550 0.4008 0.0934 0.3442 0.0711 0.3819 -0.0036 0.3594 -0.0240 0.3367 0.0083 0.3625 -0.0574 0.3961 -0.0712 0.5416 -0.0695 0.3784 … ] (1×34 double)
Sigma: [0.3112 0 0.4977 0.4414 0.5199 0.4608 0.4927 0.5207 0.5071 0.4839 0.5635 0.4948 0.6222 0.4949 0.6528 0.4584 0.6180 0.4968 0.6263 0.5191 0.6098 0.5182 0.6038 0.5275 0.5785 0.5085 0.5162 0.5500 0.5759 0.5080 … ] (1×34 double)
BoxConstraints: [351×1 double]
ConvergenceInfo: [1×1 struct]
IsSupportVector: [351×1 logical]
Solver: 'SMO'
Properties, Methods
Because the classes are inseparable, the score transformation function (ScoreSVMModel.ScoreTransform) is the sigmoid function.
Estimate scores and positive class posterior probabilities for the training data. Display the results for the first 10 observations.
[label,scores] = resubPredict(SVMModel); [~,postProbs] = resubPredict(ScoreSVMModel); table(Y(1:10),label(1:10),scores(1:10,2),postProbs(1:10,2),'VariableNames',... {'TrueLabel','PredictedLabel','Score','PosteriorProbability'})
ans=10×4 table
TrueLabel PredictedLabel Score PosteriorProbability
_________ ______________ _______ ____________________
{'g'} {'g'} 1.4862 0.82216
{'b'} {'b'} -1.0003 0.30433
{'g'} {'g'} 1.8685 0.86917
{'b'} {'b'} -2.6457 0.084171
{'g'} {'g'} 1.2807 0.79186
{'b'} {'b'} -1.4616 0.22025
{'g'} {'g'} 2.1674 0.89816
{'b'} {'b'} -5.7085 0.00501
{'g'} {'g'} 2.4798 0.92224
{'b'} {'b'} -2.7812 0.074781
Estimate the logit of posterior probabilities (classification scores) for training data using a classification generalized additive model (GAM) that contains both linear and interaction terms for predictors. Specify whether to include interaction terms when computing the classification scores.
Load the ionosphere data set. This data set has 34 predictors and 351 binary responses for radar returns, either bad ('b') or good ('g').
load ionosphereTrain a GAM using the predictors X and class labels Y. A recommended practice is to specify the class names. Specify to include the 10 most important interaction terms.
Mdl = fitcgam(X,Y,'ClassNames',{'b','g'},'Interactions',10)
Mdl =
ClassificationGAM
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'b' 'g'}
ScoreTransform: 'logit'
Intercept: 3.2565
Interactions: [10×2 double]
NumObservations: 351
Properties, Methods
Mdl is a ClassificationGAM model object.
Predict the labels using both linear and interaction terms, and then using only linear terms. To exclude interaction terms, specify 'IncludeInteractions',false. Estimate the logit of posterior probabilities by specifying the ScoreTransform property as 'none'.
Mdl.ScoreTransform = 'none'; [labels,scores] = resubPredict(Mdl); [labels_nointeraction,scores_nointeraction] = resubPredict(Mdl,'IncludeInteractions',false);
Create a table containing the true labels, predicted labels, and scores. Display the first eight rows of the table.
t = table(Y,labels,scores,labels_nointeraction,scores_nointeraction, ... 'VariableNames',{'True Labels','Predicted Labels','Scores' ... 'Predicted Labels Without Interactions','Scores Without Interactions'}); head(t)
True Labels Predicted Labels Scores Predicted Labels Without Interactions Scores Without Interactions
___________ ________________ __________________ _____________________________________ ___________________________
{'g'} {'g'} -51.628 51.628 {'g'} -47.676 47.676
{'b'} {'b'} 37.433 -37.433 {'b'} 36.435 -36.435
{'g'} {'g'} -62.061 62.061 {'g'} -58.357 58.357
{'b'} {'b'} 37.666 -37.666 {'b'} 36.297 -36.297
{'g'} {'g'} -47.361 47.361 {'g'} -43.373 43.373
{'b'} {'b'} 106.48 -106.48 {'b'} 102.43 -102.43
{'g'} {'g'} -62.665 62.665 {'g'} -58.377 58.377
{'b'} {'b'} 201.46 -201.46 {'b'} 197.84 -197.84
The predicted labels for the training data X do not vary depending on the inclusion of interaction terms, but the estimated score values are different.
Estimate in-sample posterior probabilities and misclassification costs using a naive Bayes classifier.
Load the fisheriris data set. Create X as a numeric matrix that contains four measurements for 150 irises. Create Y as a cell array of character vectors that contains the corresponding iris species.
load fisheriris X = meas; Y = species; rng('default') % For reproducibility
Train a naive Bayes classifier using the predictors X and class labels Y. A recommended practice is to specify the class names. fitcnb assumes that each predictor is conditionally and normally distributed.
Mdl = fitcnb(X,Y,'ClassNames',{'setosa','versicolor','virginica'});
Mdl is a trained ClassificationNaiveBayes classifier.
Estimate the posterior probabilities and expected misclassification costs for the training data.
[label,Posterior,MisclassCost] = resubPredict(Mdl); Mdl.ClassNames
ans = 3×1 cell
{'setosa' }
{'versicolor'}
{'virginica' }
Display the results for 10 randomly selected observations.
idx = randsample(size(X,1),10); table(Y(idx),label(idx),Posterior(idx,:),MisclassCost(idx,:),'VariableNames', ... {'TrueLabel','PredictedLabel','PosteriorProbability','MisclassificationCost'})
ans=10×4 table
TrueLabel PredictedLabel PosteriorProbability MisclassificationCost
______________ ______________ _________________________________________ ______________________________________
{'virginica' } {'virginica' } 6.2514e-269 1.1709e-09 1 1 1 1.1709e-09
{'setosa' } {'setosa' } 1 5.5339e-19 2.485e-25 5.5339e-19 1 1
{'virginica' } {'virginica' } 7.4191e-249 1.4481e-10 1 1 1 1.4481e-10
{'versicolor'} {'versicolor'} 3.4472e-62 0.99997 3.362e-05 1 3.362e-05 0.99997
{'virginica' } {'virginica' } 3.4268e-229 6.597e-09 1 1 1 6.597e-09
{'versicolor'} {'versicolor'} 6.0941e-77 0.9998 0.00019663 1 0.00019663 0.9998
{'virginica' } {'virginica' } 1.3467e-167 0.002187 0.99781 1 0.99781 0.002187
{'setosa' } {'setosa' } 1 1.5776e-15 5.7172e-24 1.5776e-15 1 1
{'virginica' } {'virginica' } 2.0116e-232 2.6206e-10 1 1 1 2.6206e-10
{'setosa' } {'setosa' } 1 1.8085e-17 1.9639e-24 1.8085e-17 1 1
The order of the columns of Posterior and MisclassCost corresponds to the order of the classes in Mdl.ClassNames.
Input Arguments
Classification machine learning model, specified as a full classification model object, as given in the following table of supported models.
| Model | Classification Model Object |
|---|---|
| Generalized additive model | ClassificationGAM |
| k-nearest neighbor model | ClassificationKNN |
| Naive Bayes model | ClassificationNaiveBayes |
| Neural network model | ClassificationNeuralNetwork |
| Support vector machine for one-class and binary classification | ClassificationSVM |
Flag to include interaction terms of the model, specified as true or
false. This argument is valid only for a generalized
additive model (GAM). That is, you can specify this argument only when
Mdl is ClassificationGAM.
The default value is true if Mdl contains interaction
terms. The value must be false if the model does not contain interaction
terms.
Data Types: logical
Output Arguments
Predicted class labels, returned as a categorical or character array, logical or numeric vector, or cell array of character vectors.
label has the same data type as the observed class labels that
trained Mdl, and its length is equal to the number of observations
in Mdl.X. (The software treats string arrays as cell arrays of character
vectors.)
Class scores, returned as a numeric matrix. Score has rows
equal to the number of observations in Mdl.X and columns equal to the
number of distinct classes in the training data
(size(Mdl.ClassNames,1)).
Expected misclassification costs, returned as a numeric matrix. This output applies
only to k-nearest neighbor and naive Bayes models. That is,
resubPredict returns Cost only when
Mdl is ClassificationKNN or
ClassificationNaiveBayes.
Cost has rows equal to the number of observations in
Mdl.X and columns equal to the number of distinct classes in the
training data (size(Mdl.ClassNames,1)).
Cost(j,k) is the expected misclassification cost of the
observation in row j of Mdl.X predicted into class
k (in class Mdl.ClassNames(k)).
Algorithms
resubPredict computes predictions according to the corresponding
predict function of the object (Mdl). For a
model-specific description, see the predict function reference pages in
the following table.
| Model | Classification Model Object (Mdl) | predict Object Function |
|---|---|---|
| Generalized additive model | ClassificationGAM | predict |
| k-nearest neighbor model | ClassificationKNN | predict |
| Naive Bayes model | ClassificationNaiveBayes | predict |
| Neural network model | ClassificationNeuralNetwork | predict |
| Support vector machine for one-class and binary classification | ClassificationSVM | predict |
Extended Capabilities
Usage notes and limitations:
This function fully supports GPU arrays for a trained classification model specified as a
ClassificationKNN,ClassificationNeuralNetwork, orClassificationSVMobject.
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2012aresubPredict fully supports GPU arrays for ClassificationNeuralNetwork.
Starting in R2023b, the following classification model object functions use observations with missing predictor values as part of resubstitution ("resub") and cross-validation ("kfold") computations for classification edges, losses, margins, and predictions.
In previous releases, the software omitted observations with missing predictor values from the resubstitution and cross-validation computations.
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)